XCMS Online软件
一、产品背景
XCMS plus是基于XCMS和XCMS在线平台设计的非靶向代谢组学,是被引用最多的代谢组学软件,用于寻找和识别代谢物。它能提供液质数据的保留时间对齐,特征检测和匹配,以及统计分析的功能。它包含超过60万分析物的质谱数据库,并拥有批处理功能,可实现最常归的数据可视化呈现。此外,XCMS plus还通过与METLIN串联质谱和METLIN中性丢失数据库集成,可以进行更准确的识别。
二、产品介绍

三、产品操作步骤

1.0 创建用户帐户
Creating a User Account
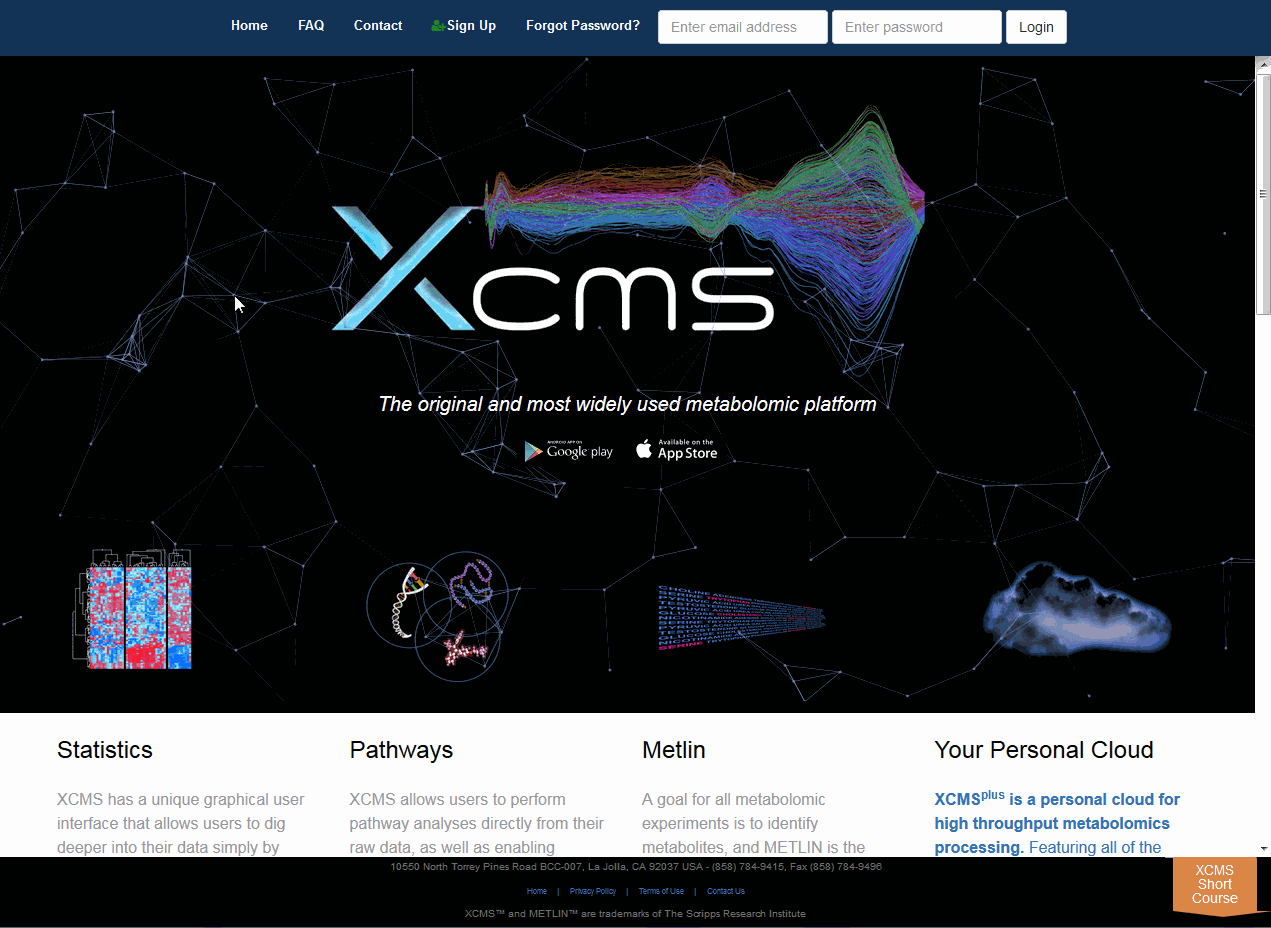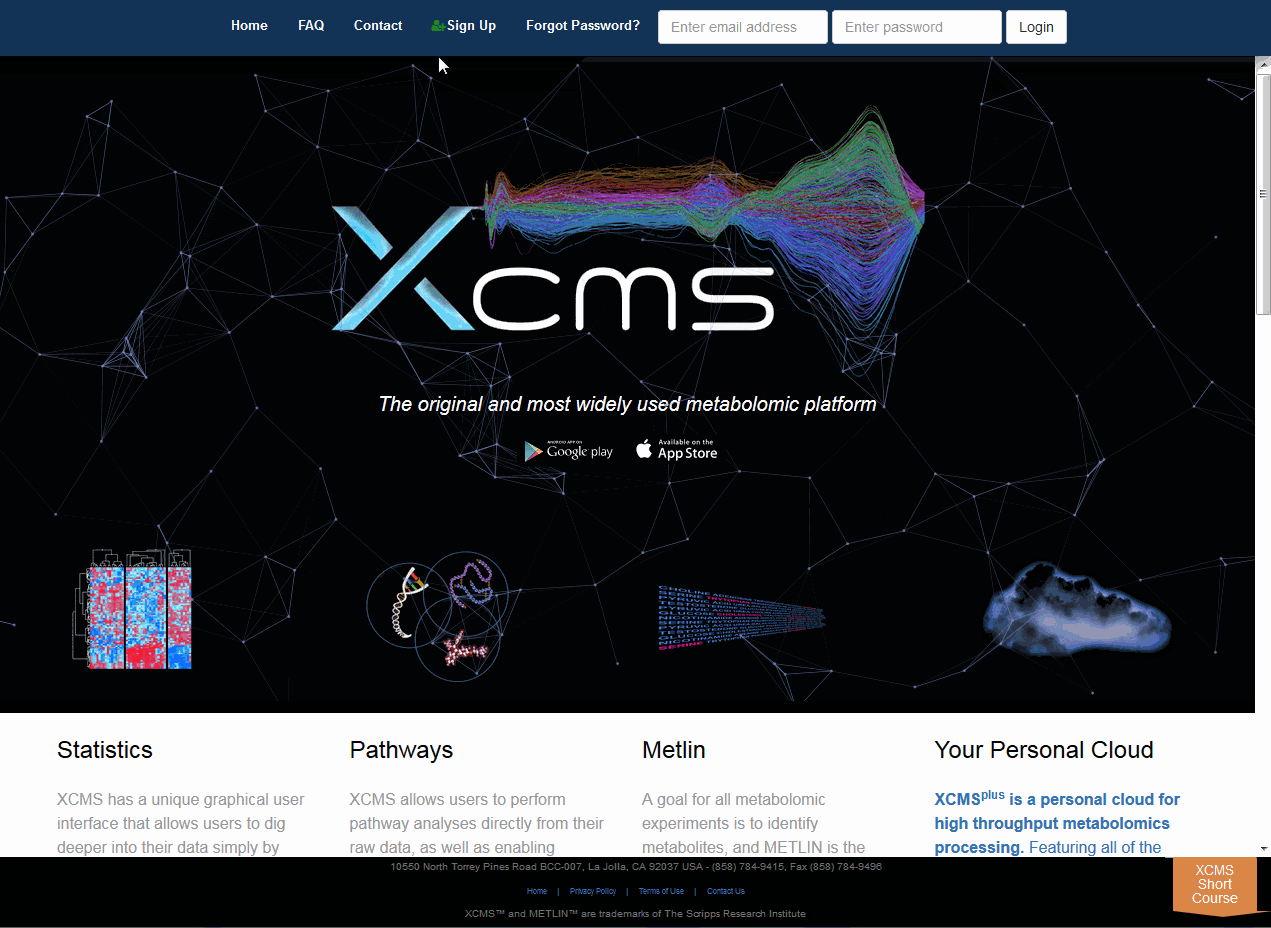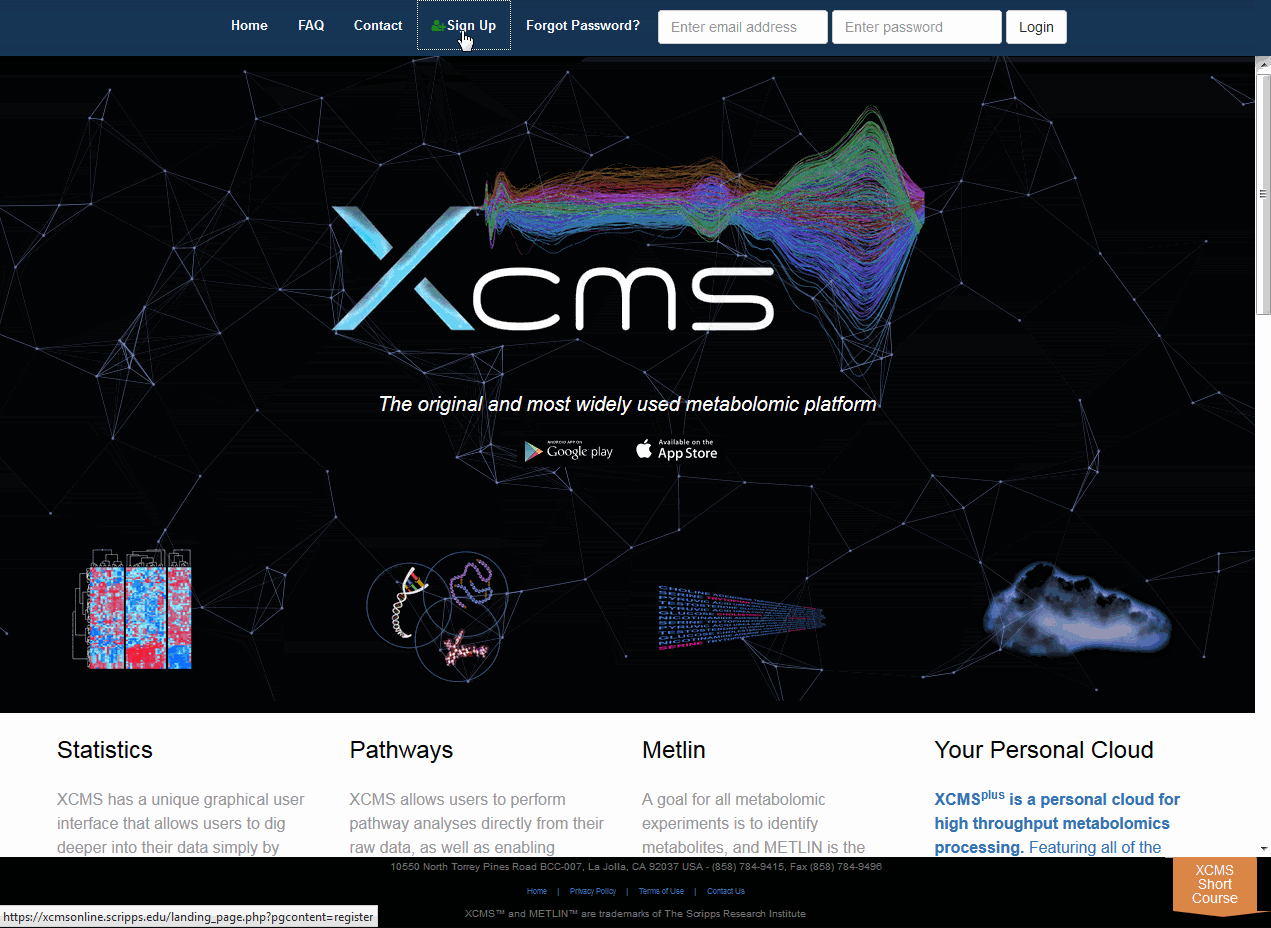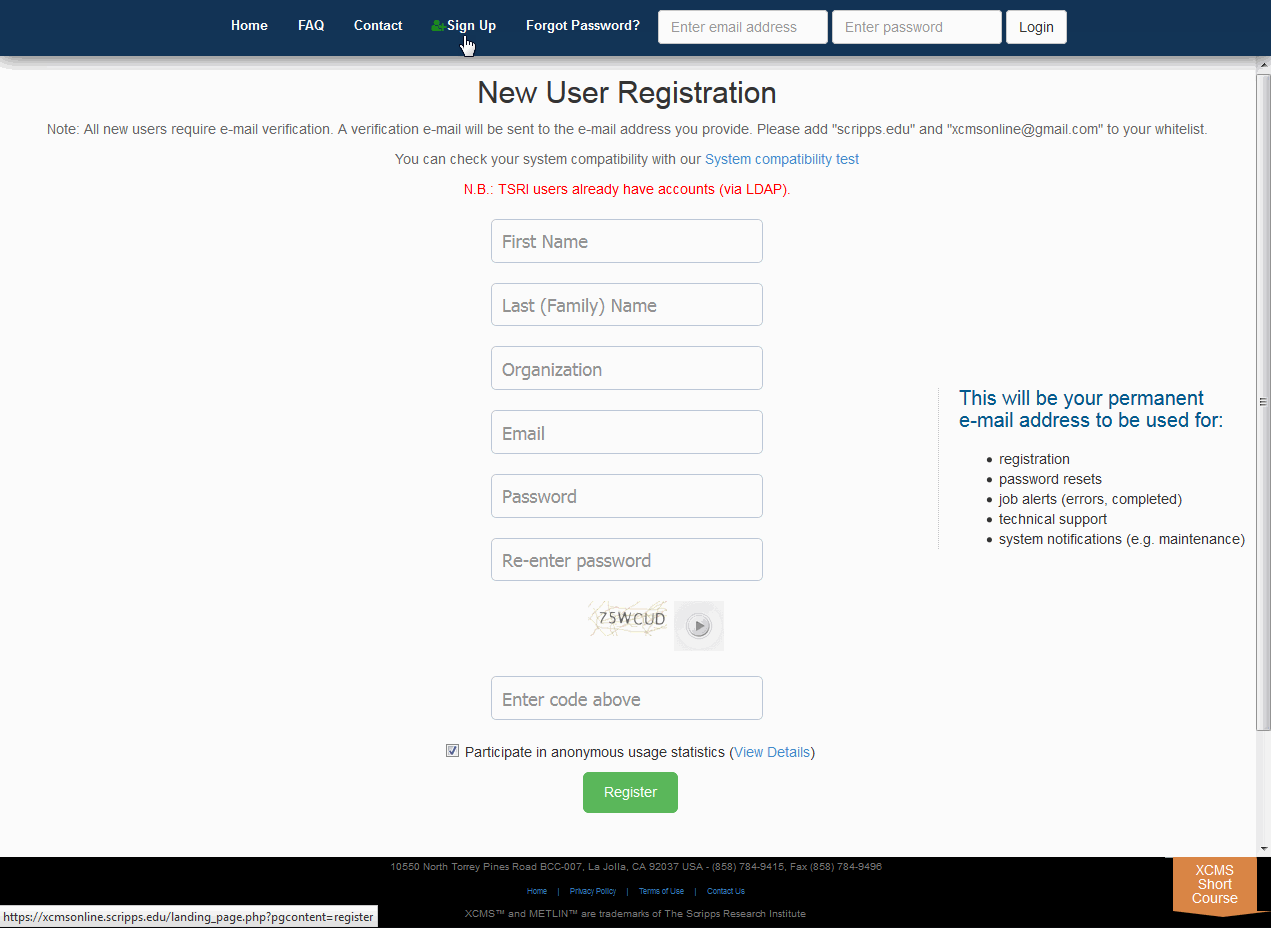
You will initially need to register your e-mail account to be able to use the system. Some configurations may already have user accounts for you or the system may authenticate against a pre-existing database such as Active Directory/LDAP. Check with your system administrator more information. Click "Sign Up" from the top navigation menu then fill out all the fields and click the "Register" button. You will receive an e-mail at the account you entered with validation instructions. Upon validation you will have access to the system. You may then login using the user name and password combination you used to register.
您首先需要注册您的电子邮件帐户才能使用系统。某些配置可能已经为您提供了用户帐户,或者系统可能根据预先存在的数据库(如Active Directory/LDAP)进行身份验证。请向系统管理员查询更多信息。点击顶部导航菜单中的“注册”,然后填写所有字段,点击“注册”按钮。您将在您输入的帐户收到一封电子邮件,里面有验证说明。验证后,您将可以访问系统。然后,您可以使用注册时使用的用户名和密码组合登录。
2.0 数据分析
Data Analysis
XCMS Online breaks data analysis up into four categories. Each category uses similar data upload system and generally follows the same step-by-step procedure: choose a job type, upload or choose dataset(s), define parameters, and submit the job. The job types can be broken up into the follow systems, each serving a different purpose:
XCMSOnline将数据分析分为四类。每个类别都使用类似的数据上传系统,通常遵循相同的步骤:选择作业类型、上载或选择数据集、定义参数,并提交作业。作业类型可以分为以下系统,每个系统具有不同的用途:
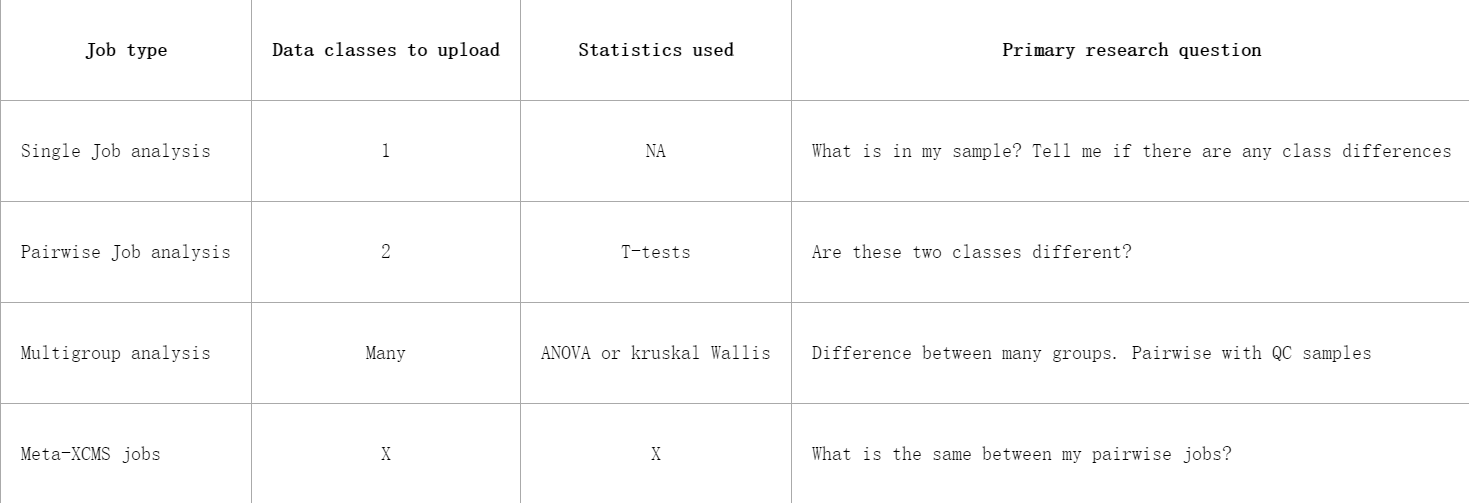
All of these data analysis types run peak detection, grouping of peaks across files into features, an alignment of features that have drifted, filling in intensity values from peaks that were missing from samples.
所有这些数据分析类型都运行峰值检测,将文件中的峰值分组为特征,对齐漂移的特征,填充样本中缺失的峰值的强度值。
2.1 数据上传准备
Data Upload Preparation
Before running an experiment you must have data in the proper format. XCMS supports the following datatypes:
mzXML、mzData、mzData.XML、netCDF、cdf、wiff、wiff.scan
There are two different uploading tools, one written in HTML5 (default) and the other in Java. The Java uploader is now considered a legacy item and will not be supported after May 2016. Also, depending on your security policy you may not be able to use Java through a browser.
NOTE: In addition the the files above, XCMS also supports folders with “.d” extension (Agilent/Bruker). You can upload these folders directly with the Java uploader, but not the HTML5 tool. To upload this data with the HTML5 tool, place the folder contents into a .ZIP archive and upload that file.
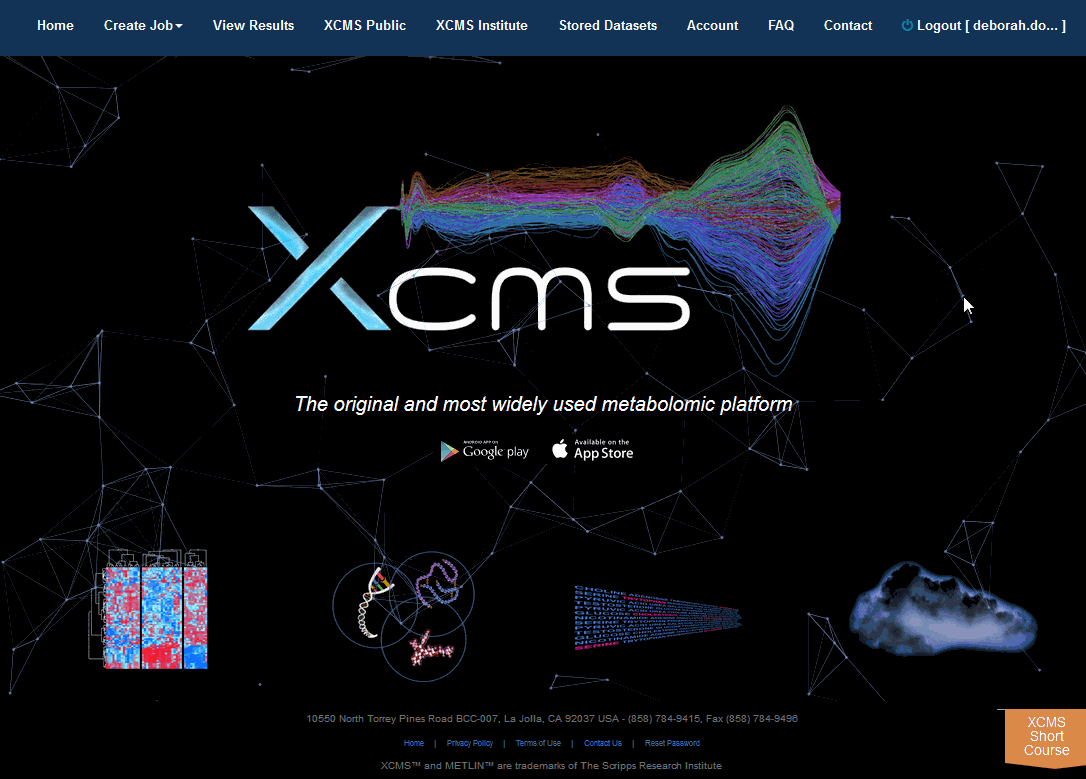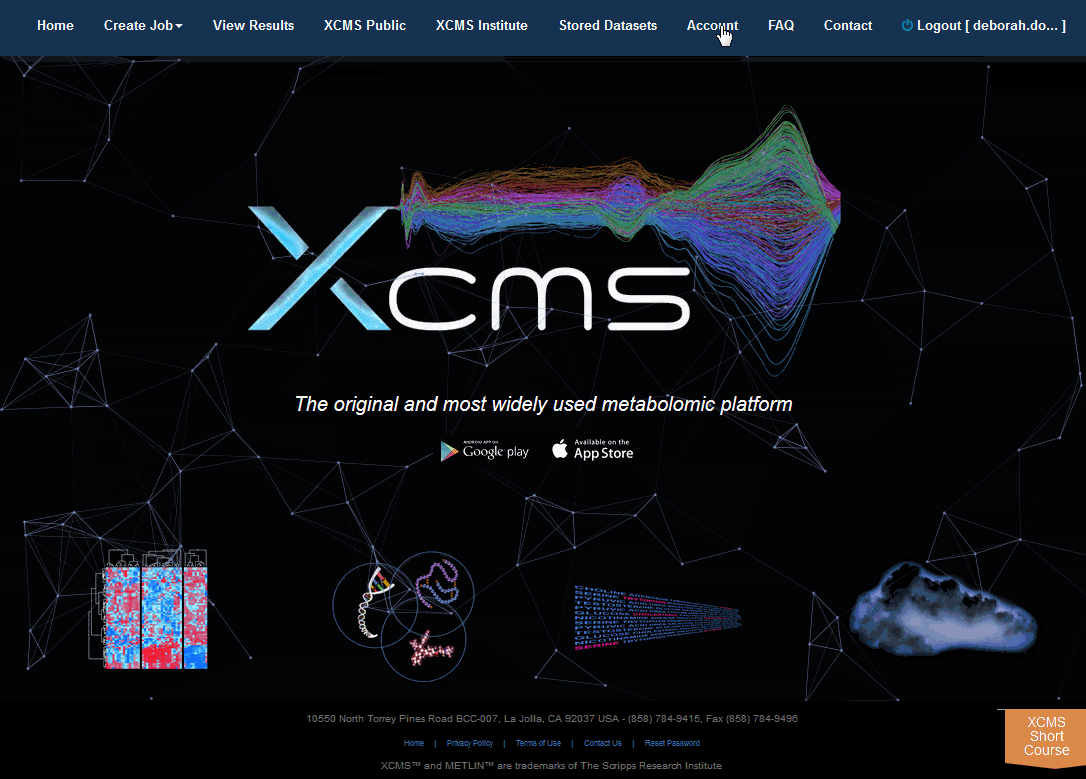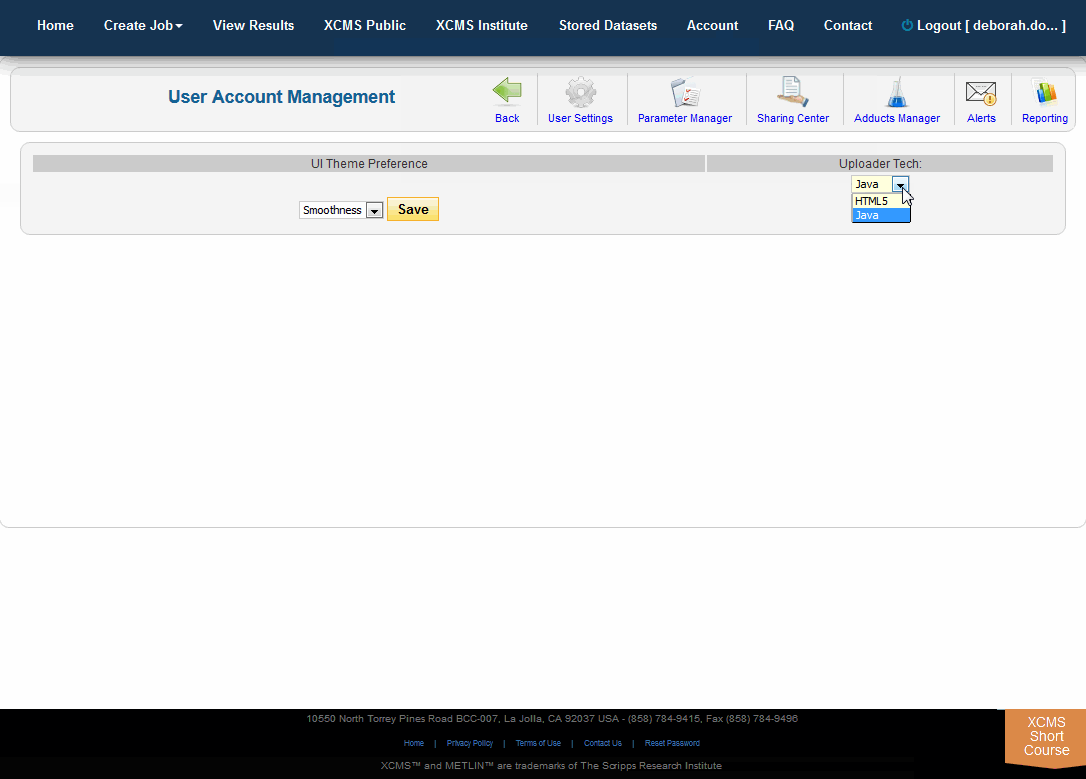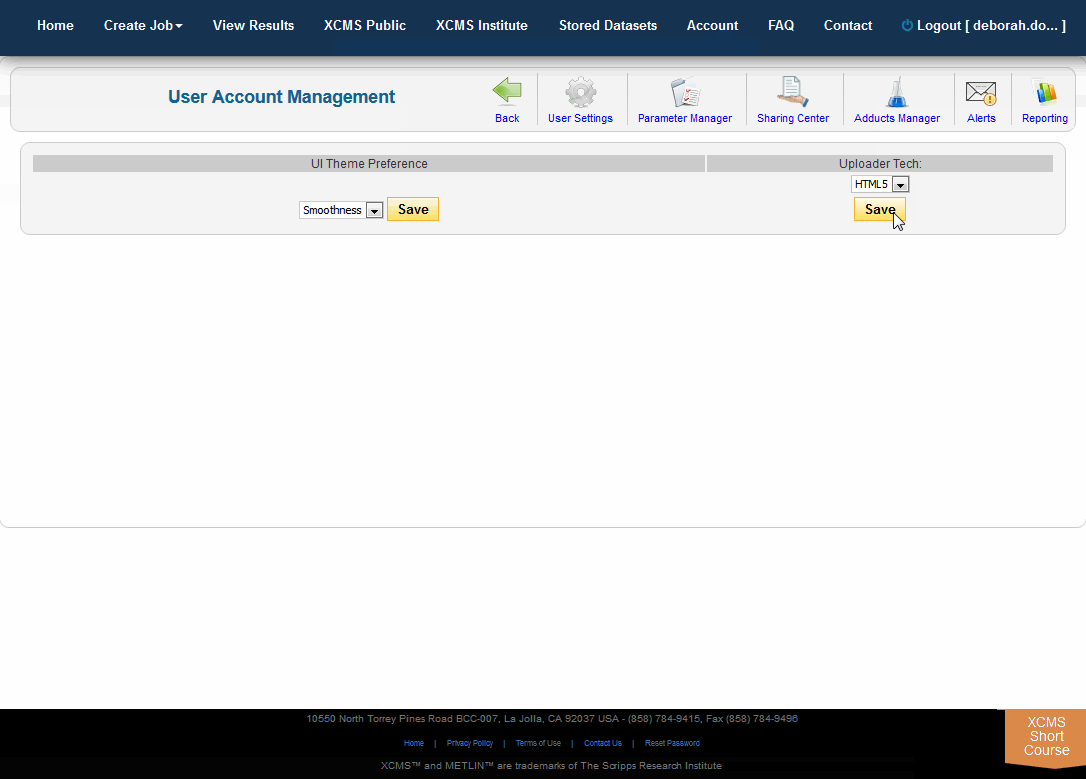
To change your default uploader, choose "Account" from the top navigation menu then "User Settings." On the right side under "Uploader Tech" choose HTML5 from the drop-down menu and click the "Save" button.
在进行实验之前,您必须有正确格式的数据。XCMS支持以下数据类型: mzXML、mzData、mzData.XML、netCDF、Cdf、wiff、wiff.scan。有两种不同的上传工具,一种用HTML5(默认)编写,另一种用Java编写。Java上传器现在被认为是一个遗留项目,在2016年5月之后将不被支持。此外,根据您的安全策略,您可能无法通过浏览器使用Java。
注意:除了上述文件外,XCMS还支持扩展名为“。d”的文件夹(安捷伦/Bruker)的文件夹。您可以直接使用Java上传器上传这些文件夹,但不能使用HTML5工具。要使用HTML5工具上载此 数据,请将文件夹内容放入a。压缩并存档然后上载该文件。
要更改您的默认上传程序,在顶部导航菜单中选择“帐户”,然后选择“用户设置”。在右侧的“Uploader Tech”下拉菜单中选择HTML5,点击“Save”按钮。
2.2 上传数据集
Uploading Datasets
The first step for running an experiment usually involves uploading data. Choosing Single, Pairwise, or Multigroup from the "Create Job" sub-menu will give you a step-by-step wizard. The fist step is to Select Datasets. Choosing "Load New Dataset" will pop-up a window with the uploader of your choice. When uploading data keep these important points in mind:
运行实验的第一步通常包括上传数据。从“Create Job”子菜单中选择单一、成对或多组,将提供一个循序渐进的向导。第一步是选择数据集。选择“Load New Dataset”将弹出一个窗口,显示您所选择的上传程序。当上传数据时,请牢记以下要点:
Dataset names must be unique. The upload tool will generate a generic name based on your username and the current date. You can change the name while the data is uploading or after it is complete.
The upload tool must remain open while data is uploading. If you close the window you must start over. However, you may minimize the window and continue working while data is uploading.
The HTML5 upload tool allows users to drag and drop files into the area labeled "Drop Here." each file has a circular progress indicator. Clicking the icon to the right of the indicator allows you to cancel the upload of or delete that file from the dataset. You can also rename the dataset at this time. When the upload is complete, click the "Save Dataset & Proceed" button at the top of the window.
数据集名称必须为唯一的名称。上载工具将根据您的用户名和当前日期生成一个通用名称。您可以在数据上载时或在数据完成后更改名称。
上传数据时,上传工具必须保持打开状态。如果你关闭了窗口,则必须重新开始。但是,您可以最小化窗口并在数据上传时继续工作。
HTML5上传工具允许用户将文件拖放到“drop Here”区域,每个文件都有一个循环进度指示器。单击指示器右侧的图标可以取消上传或从数据集中删除该文件。此时还可以重命名数据集。当上传完成后,点击窗口顶部的“Save Dataset & Proceed”按钮。
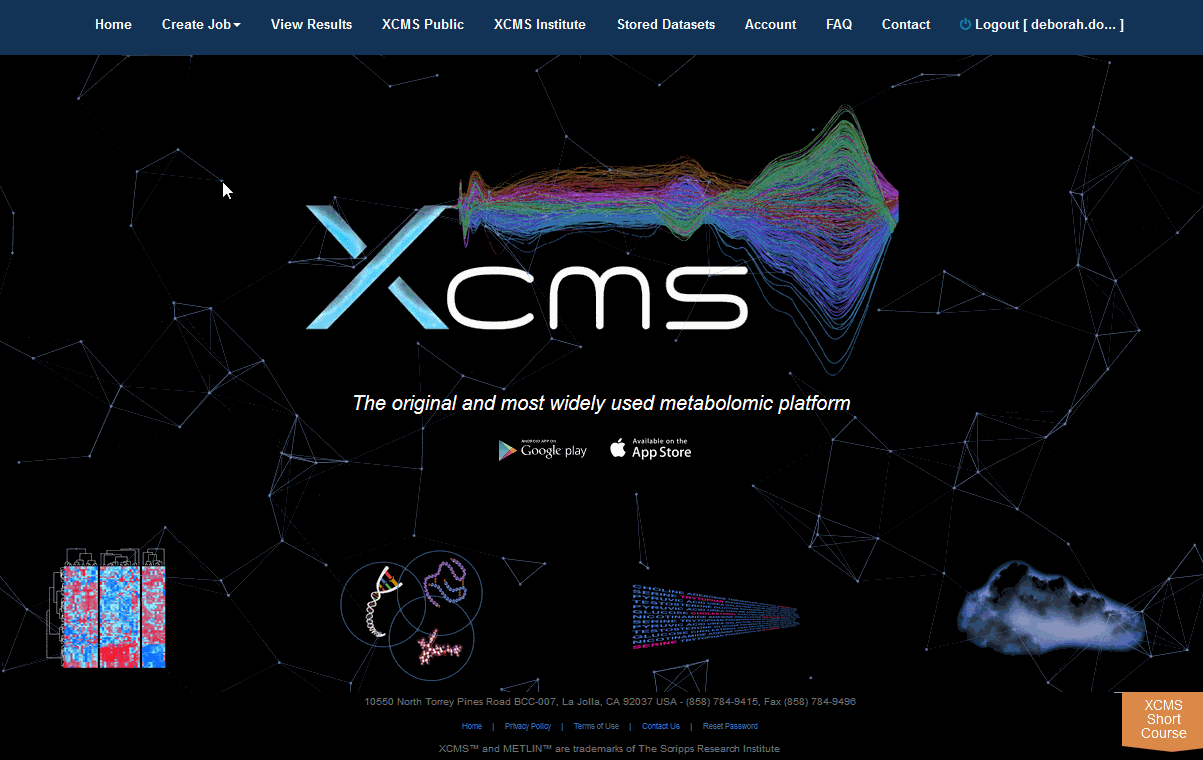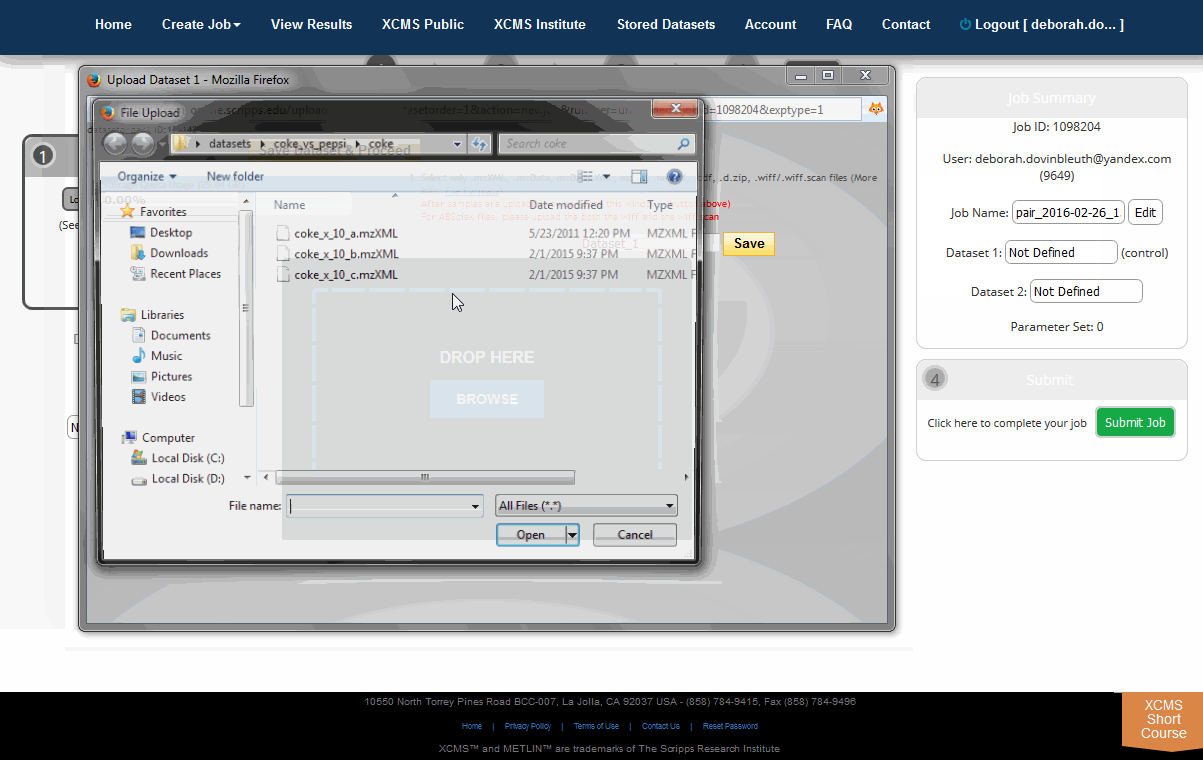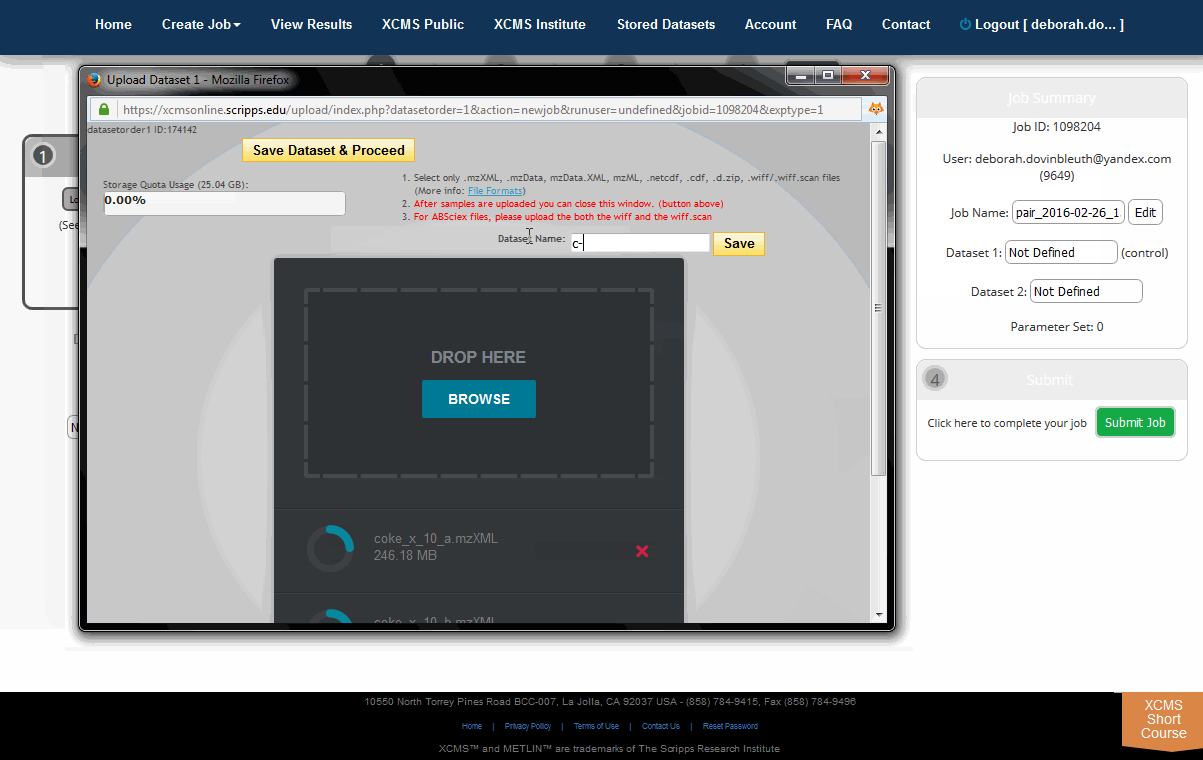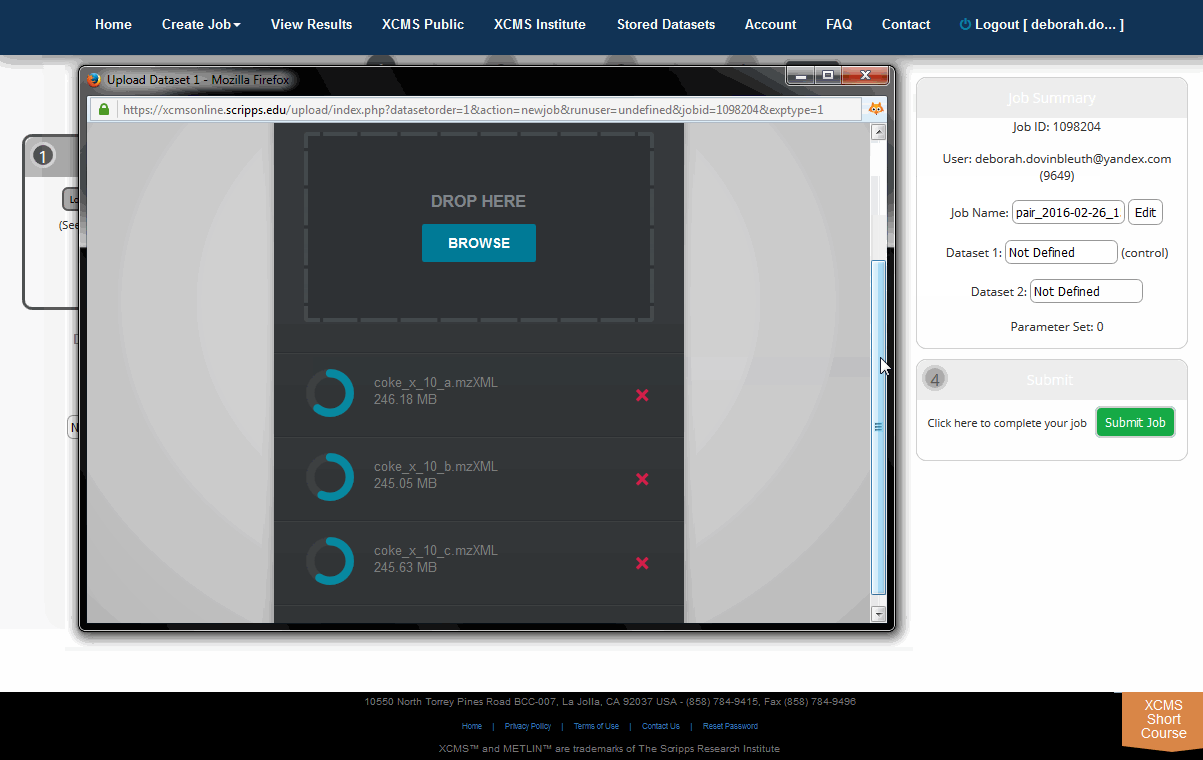
The Java upload tool may require additional setup depending on your security settings. Appendix I: Java Issues shows a few common dialog boxes you may encounter. If you receive a warning, click “Run” or add an exception for this server. Once the Java applet has permission to run you will see an upload dialog box.
Java上传工具可能需要额外的设置,这取决于您的安全设置。附录I: Java问题显示了您可能会遇到的一些常见对话框。如果收到警告,单击“运行”或为该服务器添加一个例外。一旦Java applet拥有运行权限,您将看到一个上传对话框。
Select files/folders to be included in the dataset from the file tree. Individual files will appear in the file list as you navigate the file tree. After you locate the files for this dataset, drag them to the “Drop Files Here” area located in the lower-right of the screen and they will immediately start compressing (zip compression). Alternatively you can click the “+” symbol to select the files you want to include. You may need to maximize window size or use scroll bars to display all text. A small progress bar will appear under each file as it is zipped. You may need to wait until zipping finishes before clicking the "Upload" button. When you have included all the necessary files, press the Upload button and move on the the next step.
从文件树中选择要包含在数据集中的文件/文件夹。在文件树中导航时,单个文件将出现在文件列表中。定位到此数据集的文件后,将它们拖到屏幕右下角的“Drop files Here”区域,它们将立即开始压缩(zip压缩)。或者,你可以点击“+”符号来选择你想要包含的文件。您可能需要最大化窗口大小或使用滚动条来显示所有文本。当压缩文件时,每个文件下都会出现一个小的进度条。您可能需要等到压缩完成后再点击“上传”按钮。当您包含了所有必要的文件后,按下Upload按钮并进入下一步。
2.3 成对组分分析
Pairwise Job Creation
The most common experimental design in metabolomics is two-group analysis, where “disease” and “control” or “before” and “after” treatment groups are compared. Even in a simple two-group experiment, choosing the right statistical test may be a challenging task for users without a background in the field of biostatistics. Depending on data distribution and experimental design, XCMS Online offers the choice of parametric or nonparametric, independent (unpaired) or dependent (paired) two-group tests. In general, two-group tests allow users to determine the metabolite features whose levels are significantly different between two defined conditions.
代谢组学中最常见的实验设计是两组分析,将“疾病”和“对照组”或“治疗前”和“治疗后”组进行比较。即使是在一个简单的两组实验中,对于没有生物统计学背景的用户来说,选择正确的统计测试可能也是一项具有挑战性的任务。根据数据分布和实验设计,XCMS Online提供了参数或非参数、独立(未配对)或依赖(配对)两组检验的选择。一般来说,两组测试允许用户确定代谢物的特征,其水平在两种确定的条件下显著不同。
Step 1: Create a Job. Click on “Create Job” in the top navigat ion menu. If you do not select an experiment type, the system defaults to pairwise.
Step 2: Upload the Datasets. The create job page will open for you to define the job. There is a step navigation wizard directly below the top navigation bar that will serve as a guide. Upon initial load, a default job name will be created based on your user name and the current date. Clicking the job name allows you to edit it.
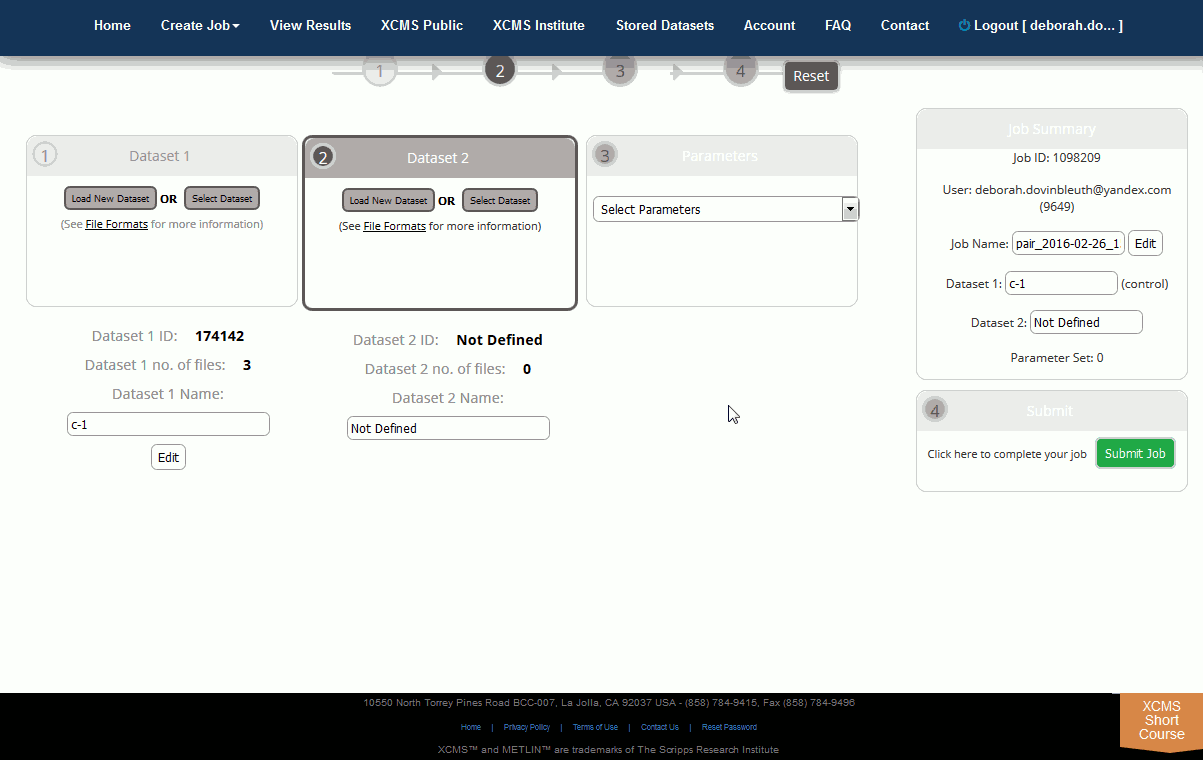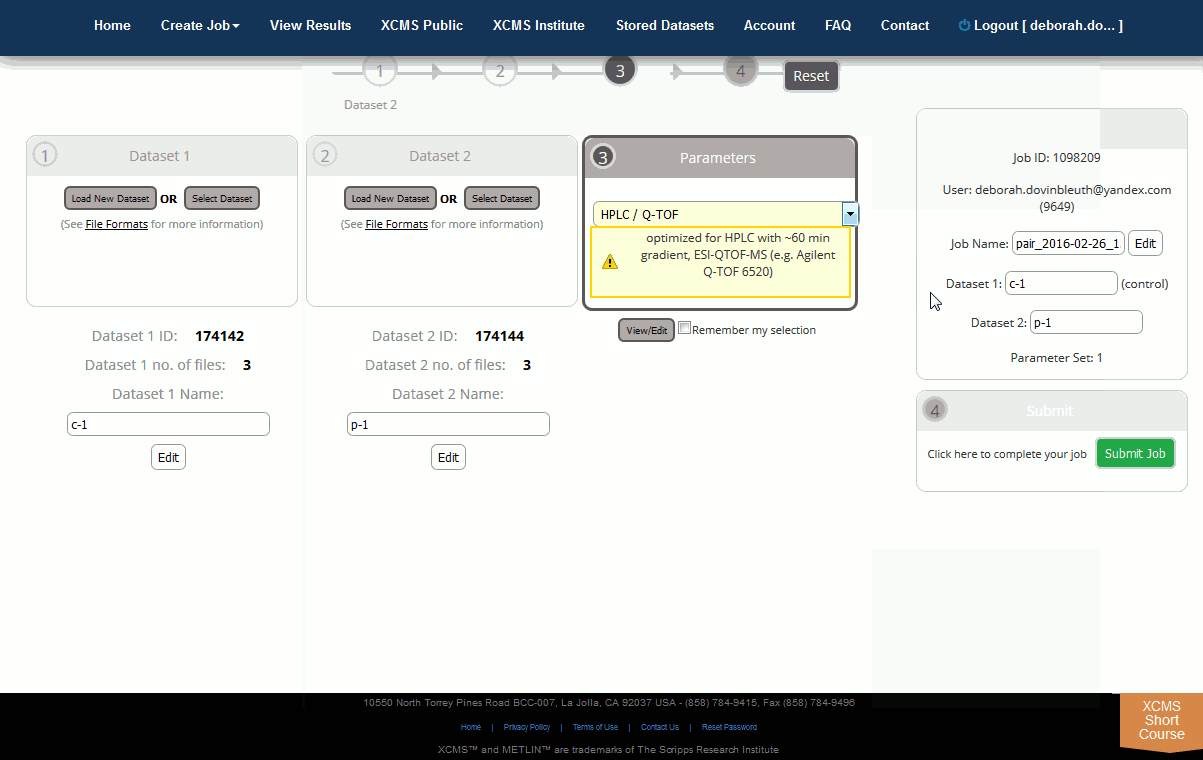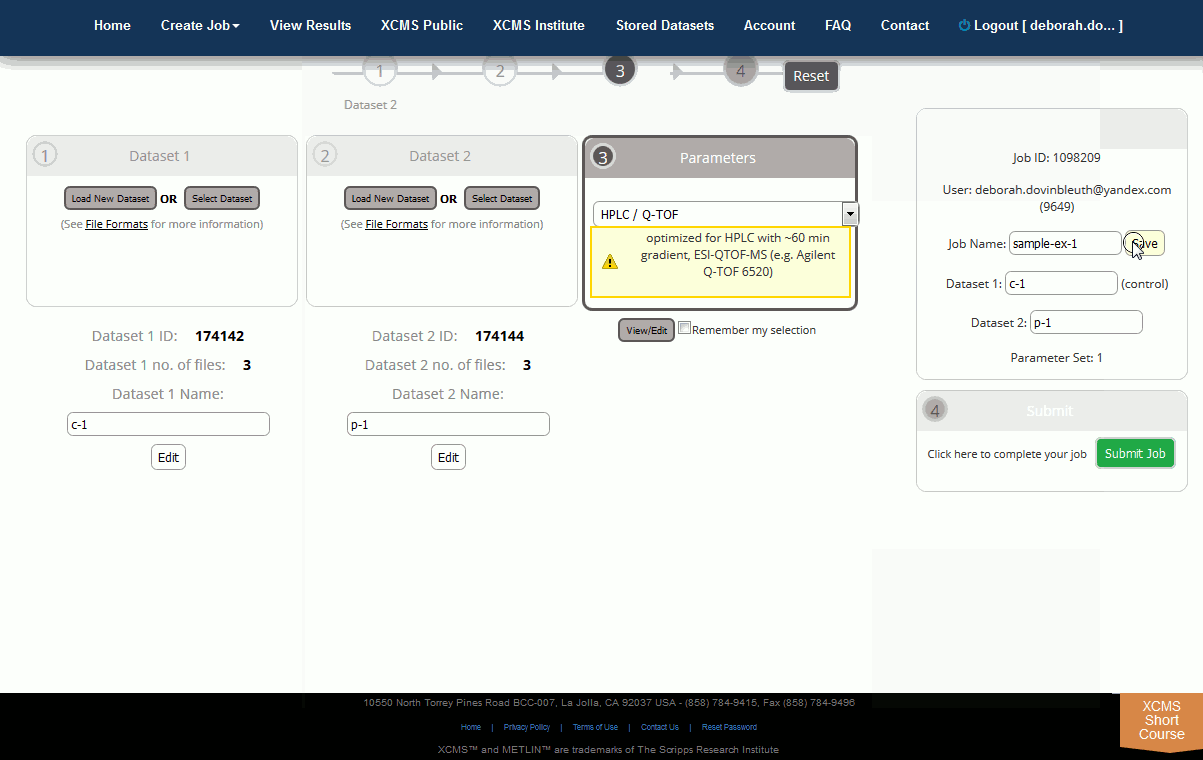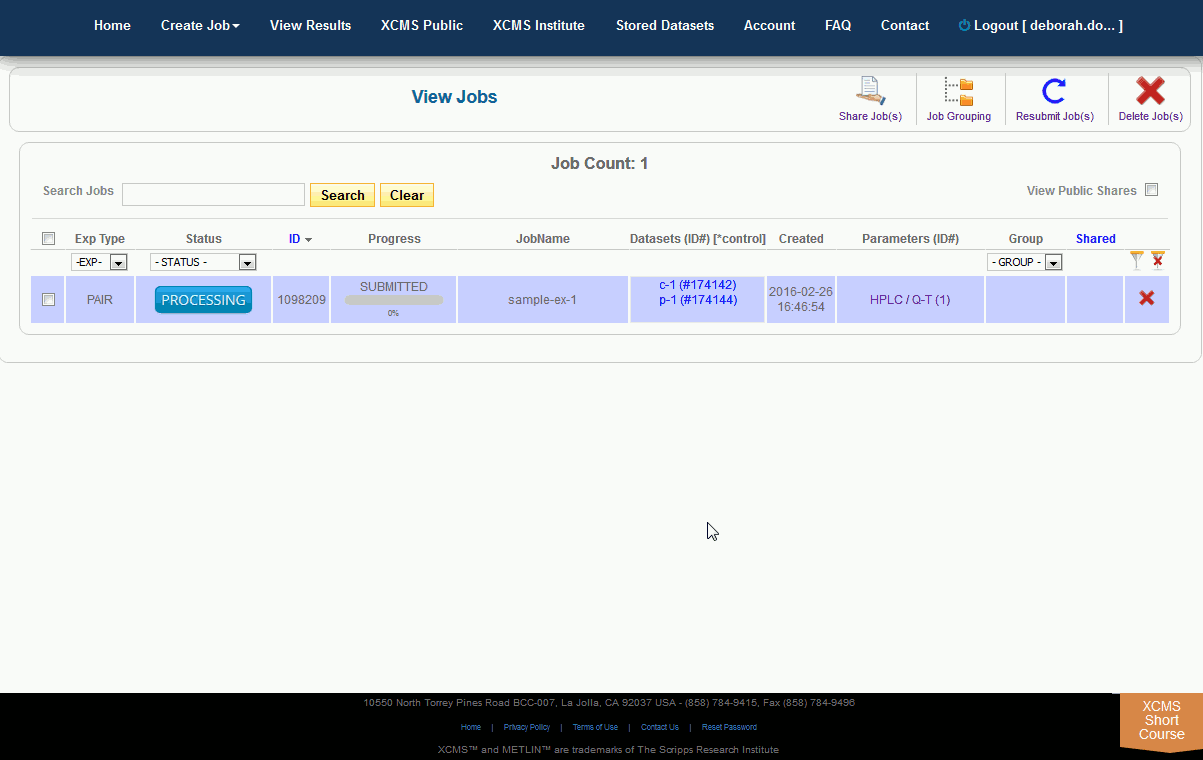
Initially you will need to upload a dataset. Dataset 1 is often defined as the control data set in pairwise jobs (left-hand side). After this is completed you can begin the second dataset upload. The Dataset 2 section can be found to the right of the Dataset 1 section. Remember, you must have unique names for each dataset even if the ID is different. You can modify the dataset names after an upload has started by clicking the “Edit” button to the right of the respective dataset name.
Step 3: Select Parameter Set. After uploads are defined you must define the corresponding job parameters. This can be done while uploads are still in progress. For your convenience XCMX Online offers predefined parameter set templates (e.g. HPLC/TOF, UPLC/TOF, HPLC/OrbiTrap). Simply select one and customize it to your particular needs. Click on the Parameter drop down box to begin defining parameters. After you select a template, the dropdown will change and you will be able to view or edit the selected parameter set.
Step 4: Define Custom Parameters. Skip this step if you do not wish to define custom parameters. Select the “View/Edit” button. This will present a modal dialog box where you can define how the datasets are processed. Methods and properties are available for all aspects of dataset processing.
If you choose to change any of the settings in a template parameter set, you will need to create a new parameter name for processing otherwise your changes will not be saved. You cannot overwrite a template parameter set. The name for a custom parameter set can be set in the “General” tab.
Step 5: Submit the Job. At this point you can review all relevant details about the job prior to submission (e.g. job name, dataset names, correct file count, correct parameters). Once you are satisfied, click the “Submit Job” button to queue the job for processing. The “Submit Job” button can be found at the bottom of the screen. Even if you still have datasets uploading you can pre-submit the job and it will automatically queue once the upload is complete.
第 1 步:创建作业。单击顶部导航菜单中的"创建作业"。如果您没有选择实验类型,系统将默认为对。
第 2 步:上传数据集。创建作业页面将打开,供您定义作业。顶部导航栏正下方有一个步骤导航向导,用作向导。初始加载后,将根据您的用户名和当前日期创建默认作业名称。单击工作名称可以编辑它。最初,您需要上传数据集。数据集 1 通常定义为对齐作业中的控制数据集(左侧)。完成此问题后,您可以开始第二个数据集上传。数据集 2 部分位于数据集 1 部分的右侧。请记住,即使 ID 不同,您也必须为每个数据集提供唯一的名称。上传开始后,您可以单击相应数据集名称右侧的"编辑"按钮来修改数据集名称。
第 3 步:选择参数集。定义上传后,您必须定义相应的工作参数。这可以在上传仍在进行中时完成。为了方便起见,XCMS Online提供预先定义的参数设置模板(例如 HPLC/TOF、UPLC/TOF、HPLC/奥比陷阱)。只需选择一个,并根据您的特定需求自定义它。单击参数下拉框开始定义参数。选择模板后,下拉将发生变化,您将能够查看或编辑所选参数集。
第 4 步:定义自定义参数。如果您不想定义自定义参数,请跳过此步骤。选择"查看/编辑"按钮。这将呈现一个模态对话框,您可以定义如何处理数据集。数据集处理的所有方面都提供方法和属性。如果您选择更改模板参数集中的任何设置,则需要创建一个新的参数名称进行处理,否则将无法保存您的更改。您不能覆盖模板参数集。自定义参数集的名称可以设置在"一般"选项卡中。
第 5 步:提交作业。此时,您可以在提交之前查看有关作业的所有相关详细信息(例如工作名称、数据集名称、正确的文件计数、正确的参数)。满意后,单击"提交作业"按钮以排队处理作业。"提交作业"按钮可在屏幕底部找到。即使您仍有上传数据集,您也可以预提交作业,并在上传完成后自动排队。
After you click “Submit Job”, you will have a final opportunity to view all settings for the job prior to submission. A modal dialog box will appear on your screen displaying job settings. Carefully review the information as this cannot be changed after submission (You would need to create a new job, although your data sets will already be stored). If you are satisfied with the displayed information, click the “Submit Job” button in the lower-right corner of the dialog box, otherwise click cancel and make the necessary changes.
单击"提交作业"后,您将有最后的机会在提交之前查看作业的所有设置。显示工作设置的屏幕上将显示模态对话框。仔细审查信息,因为提交后无法更改此信息(您需要创建新工作,尽管您的数据集已经存储)。如果您对显示的信息感到满意,请单击对话框右下角的"提交作业"按钮,否则单击"取消并进行必要的更改"。
It is important to not close the browser if you have uploads in progress as these will cancel. There is a timer on the page that will automatically redirect the entire page to the “View Results” section when all uploads are complete. XCMS Online will notify you when your job has successfully queued.
重要的是,如果有正在上传的不要关闭浏览器,因为这些会被取消。页面上有一个定时器,当所有上传完成时,该定时器将自动将整个页面重定向到"查看结果"部分。XCMS Online将在您的工作成功排队时通知您。
2.4 单组分分析
Single Job Creation
The single job analysis is best suited to the analysis when either the researcher has no known class description for their data. With this analysis there are no univariate statistics so the job does not return p-values for each feature. However, multivariate statistics such as principal component analysis will be available and will help the user decide if any classes/grouping is occurring in the dataset. To do single job analysis perform the following tasks:
单个作业分析最适合分析时,要么研究人员没有已知的类描述,他们的数据。通过此分析,没有一次性的统计数据,因此工作不会返回每个功能的 p 值。但是,将提供多变量统计(如主要组件分析),并帮助用户决定是否在数据集中出现任何类/分组。执行单一工作分析执行以下任务:
Step 1: Create a Job. This will open the interface for the creation of a single job type.
Step 2: Select Dataset(s). Choose either upload data or select an existing dataset. For uploading a new dataset see section 3.2. only one dataset should be uploaded/selected. The dataset can contain many files or just one file.
Step 3: Select Parameters. Select a preset or customized parameter set for processing your job. For details on parameter select please see section 7.0.
Step 4: Submit the Job. Check that the options for the job are as you require and optionally change the name of the job. Finally, if everything looks correct click submit.
第 1 步:创建作业。这将打开创建单个工作类型的界面。
第 2 步:选择数据集。选择上传数据 或选择现有数据集。有关上传新数据集,请参阅第3.2节。只应上传/选择一个数据集。数据集可以包含许多文件或仅包含一个文件。
第 3 步:选择参数。选择用于处理作业的预设或自定义参数集。有关参数的详细信息,请参阅第7.0节。
第 4 步:提交作业。检查作业的选项是否根据您需要,并选择更改作业名称。最后,如果一切看起来都是正确的,请单击提交。
2.5 多组分分析
Multigroup Job Creation
Multiple group analysis is an extension of two-group analysis that allows the comparison of means for multiple independent groups and enables the identification of metabolite features whose variation pattern is statistically significant. To evaluate the metabolite variation across different experimental groups, XCMS Online provides the univariate analysis of variance (one-way ANOVA) as a parametric test option and the Kruskal−Wallis test as its nonparametric alternative. The post-hoc analysis is used to determine which groups significantly differ in their metabolite expression pattern. Multigroup cloud plots display the metabolite features whose level varies significantly across different analyzed groups or data classes.
多组分析是两组分析的延伸,允许比较多个独立组的手段,并能够识别其变异模式具有统计学意义的代谢物特征。为了评估不同实验组的代谢物变异,XCMS Online 将方差 (单向 ANOVA) 作为参数测试选项,将 Kruskal −Wallis 测试作为非参数化测试选项。事后分析用于确定哪些组在代谢物表达模式上显著不同。多组云图显示代谢物特征,其水平因不同分析组或数据类别而显著不同。
Step 1: Create a Job. Begin by creating a new multigroup job. Multigroup jobs are initiated by hovering over the “Create Job” in the top navigation menu. If you do not select an experiment type, the system defaults to pairwise. You will notice there is only 1 area for adding datasets (via upload or selection) in the step navigation wizard. As you add datasets, they will show up in a list below the wizard.
Step 2: Define Biological pooled quality control (QC) dataset (OPTIONAL). This optional step allows users to select a biological pooled QC sample. This biological pooled QC currently removes this sample class from the statistical evaluation. However, the EIC and box plots of this class will still be included.
Step 3: Select Parameter Set. After uploads are defined you must define the corresponding job parameters. This can be done while uploads are still in progress. For your convenience XCMS Online offers predefined parameter set templates (e.g. HPLC/TOF, UPLC/TOF, HPLC/OrbiTrap). You can simply select one of those predefined parameter sets or customize them to your particular needs. To begin defining parameters, click on the Parameter drop down box. After you select a template, the drop-down will change and you will be able to view or edit the selected parameter set. The main difference between default multigroup and default pairwise parameter sets is the statistical tests performed (See statistics tab of parameter set for details).
Step 4: Submit the Job. At this point you can review all relevant details about the job prior to submission (e.g. job name, dataset names, correct file count, correct parameters). Once you are satisfied, click the “Submit Job” button to queue the job for processing. The “Submit Job” button can be found at the bottom-right of the screen, similar to pairwise job work-flow.
第 1 步:创建作业。首先创建新的多组作业。多组作业通过悬停在顶部导航菜单中的"创建作业"上启动。如果您没有选择实验类型,系统将默认为对。您将注意到步骤导航向导中只有 1 个区域用于添加数据集(通过上传或选择)。当您添加数据集时,它们将显示在向导下面的列表中。
第 2 步:定义生物汇集质量控制 (QC) 数据集 (可选)。此可选步骤允许用户选择生物池 QC 样本。此生物汇集 QC 当前从统计评估中删除此示例类。但是,此类的 EIC 和框图仍将包括在内。
第 3 步:选择参数集。定义上传后,您必须定义相应的工作参数。这可以在上传仍在进行中时完成。为了方便起见,XCMS Online提供预先定义的参数设置模板(例如 HPLC/TOF、UPLC/TOF、HPLC/奥比陷阱)。您可以简单地选择这些预先定义的参数集之一,或根据您的特定需求自定义它们。要开始定义参数,请单击参数下拉框。选择模板后,下拉将发生变化,您将能够查看或编辑所选参数集。默认多组和默认对向参数集之间的主要区别是执行的统计测试(有关详细信息,请参阅参数集的统计选项卡)。
第4步:提交作业。此时,您可以在提交之前查看有关该工作的所有相关细节。作业名称、数据集名称、正确的文件计数、正确的参数)。满意后,点击“提交作业”按钮,排队进行处理。屏幕右下角设有“提交作业”按钮,类似于按成对排列的作业工作流程。
2.6 meta统计分析
metaXCMS Job Creation
Metabolomic studies can reveal hundreds of dysregulated metabolic features, even using stringent statistical criteria. Meta-XCMS can be especially useful for untargeted metabolomic studies, where multiple pairwise jobs were performed and different biological sources are available.
Each meta-XCMS job requires at least two pairwise jobs each with the same control dataset. For example a meta-xcms jobs would contain a pairwise job WT vs KO, a WT vs KO2 and WT vs KO3 where each WT is from the same sample and has the same dataset ID. Once these jobs are finished users can make the meta-XCMS job as follows:
代谢研究可以揭示数百个不失调的代谢特征,即使使用严格的统计标准。Meta-XCMS对无靶向代谢酶研究特别有用,其中进行了多个成对的工作,并有不同的生物来源。每个元XCMS作业至少需要两个成对作业,每个作业都具有相同的控制数据集。例如,一个元xcms作业将包含一个成对的作业,WT和KO,WT和超氧化钾,WT和KO3,其中每个WT来自相同的样本,并具有相同的数据集ID。一旦这些作业完成,用户就可以创建meta-XCMS作业如下:
Step 1: Create a Job. You will want to begin by creating a new metaXCMS job. metaXCMS jobs are initiated by hovering over the “Create Job” in the top navigation menu. If you do not select an experiment type, the system defaults to pairwise. You can only add completed pairwise jobs as source inputs for a metaXCMS experiment so there is no uploading involved. Details about each pairwise job will appear in the table below the selection dialog as you add them.
Step 2: Select Parameter Set. Parameters for metaXCMS jobs are defined in step 2 of the wizard (center of screen). There are three (3) tabs for defining fold change, p-value, intensity cutoffs as well as alignment and adduct settings.
Step 3: Submit the Job. When you are satisfied with the parameters for the metaXCMS job, click the “Submit Job” button in the lower-right. Details about each pairwise job will appear in this table.
第1步:创建一个作业。您将希望首先创建一个新的metaXCMS作业。metaXCMS作业是通过悬停在顶部导航菜单中的“创建作业”上来启动的。如果不选择实验类型,系统默认为成对。您只能将已完成的成对工作作为metaXCMS实验的源输入,因此不涉及上载。添加每个成对作业的详细信息将显示在选择对话框下面的表中。
第2步:选择“参数集”。metaXCMS作业的参数将在向导的步骤2中定义(屏幕中心)。有三个(3)选项卡,用于定义折叠变化、p值、强度截止,以及对准和加法设置。
第3步:提交作业。当您对metaXCMS作业的参数感到满意时,请单击右下角的“提交作业”按钮。有关每个成对作业的详细信息将显示在此表中
3.0 结果
Results
After you submit your job, you will be forwarded to the “View Results” page where you can see details of your submitted jobs including progress percentage, datasets used and parameter set used. The page will automatically refresh if you have pending jobs, however, you will also receive an e-mail when each submitted job is complete. Once complete, you may view the results by clicking the green “VIEW” button. Details about each job will appear. Each experiment type will have similar visualization tools but not all will be applicable to all jobs.
提交作业后,将转发到“查看结果”页面,在那里可以查看提交作业的详细信息,包括进度百分比、使用的数据集和使用的参数集。如果您有等待的作业,该页面将自动刷新,但是,当每个提交的作业完成时,您也会收到电子邮件。完成后,您可以通过点击绿色的“查看”按钮来查看结果。此时将显示有关每个作业的详细信息。每种实验类型都有类似的可视化工具,但并非所有的都适用于所有作业。
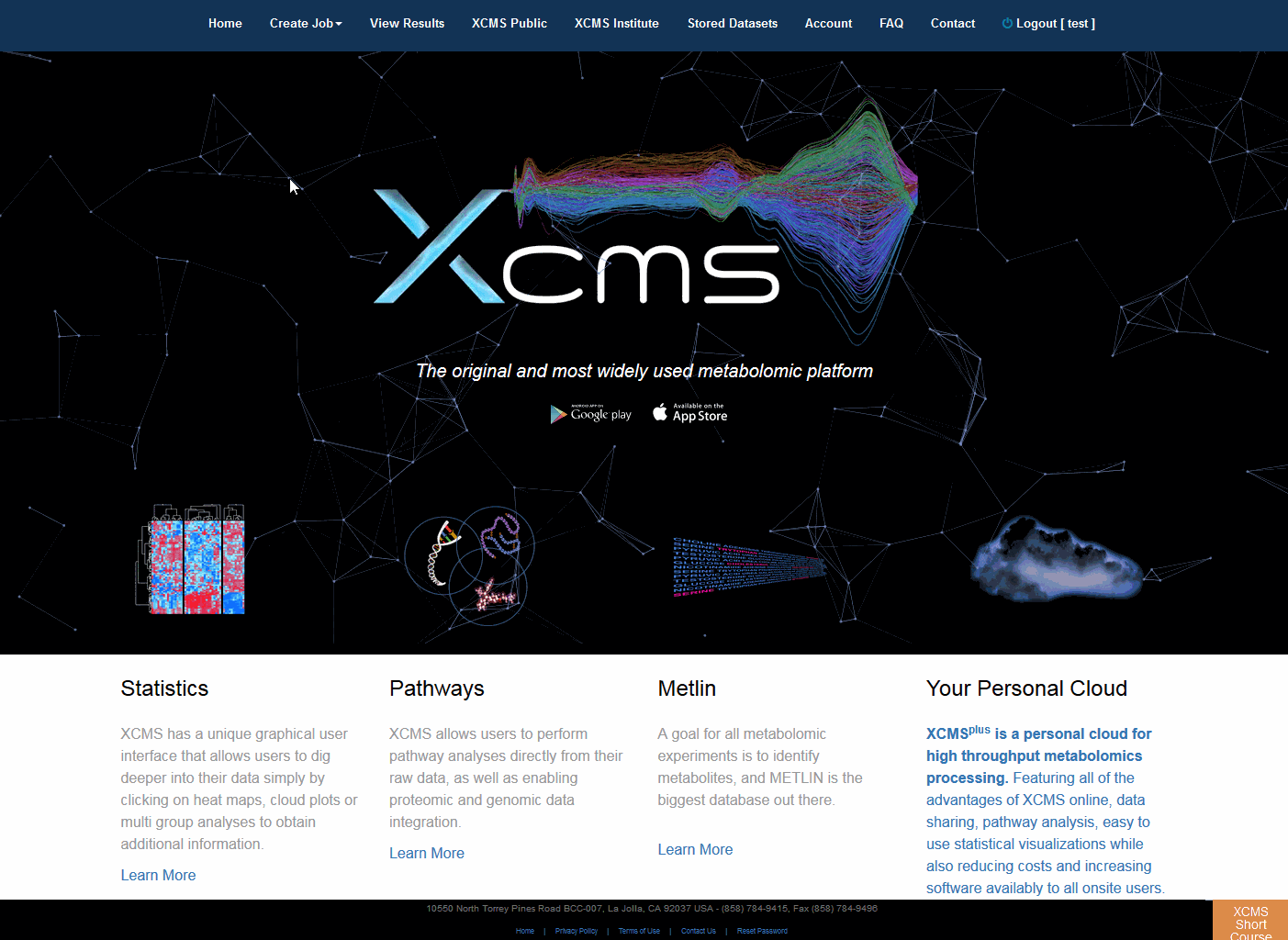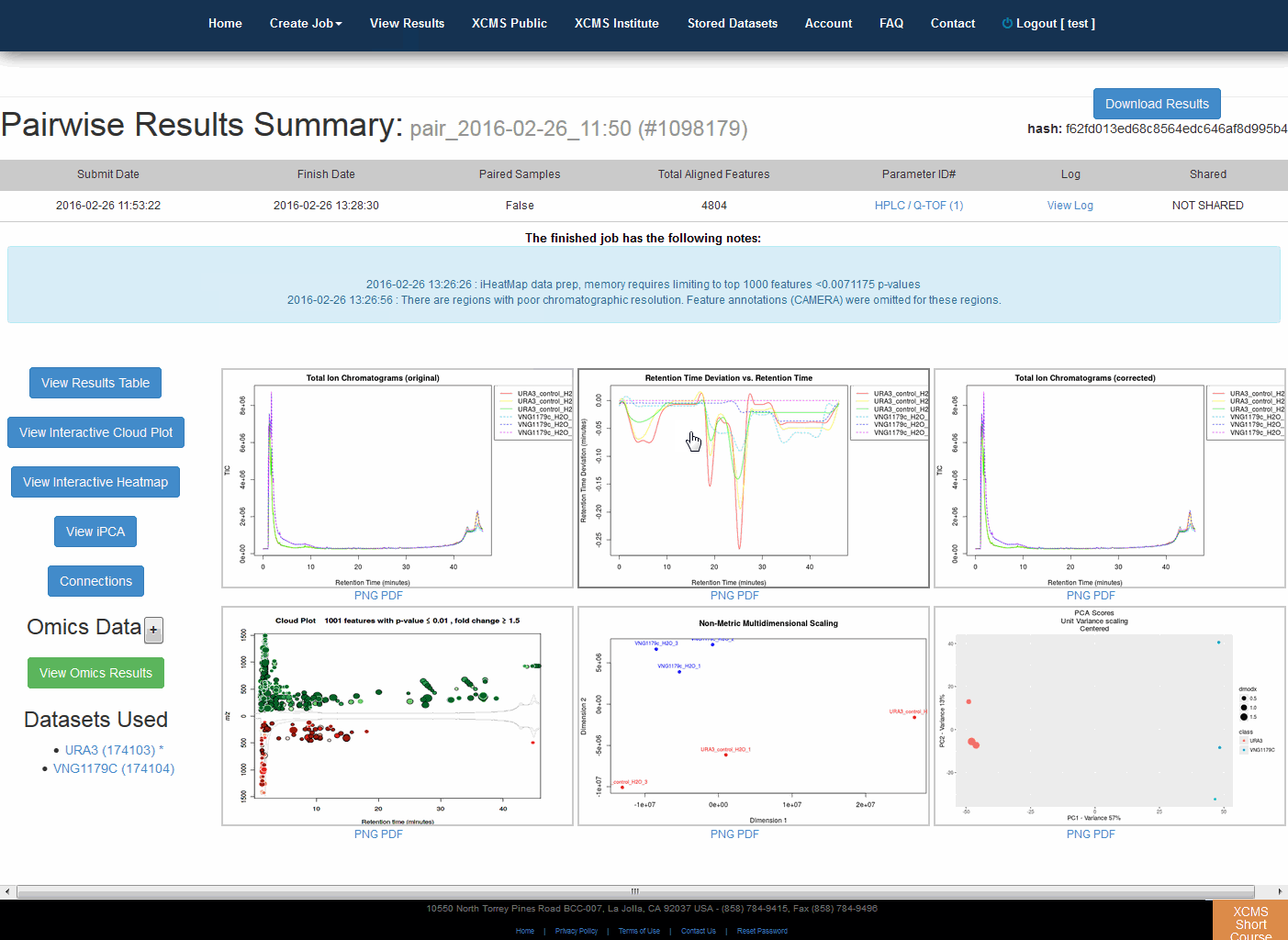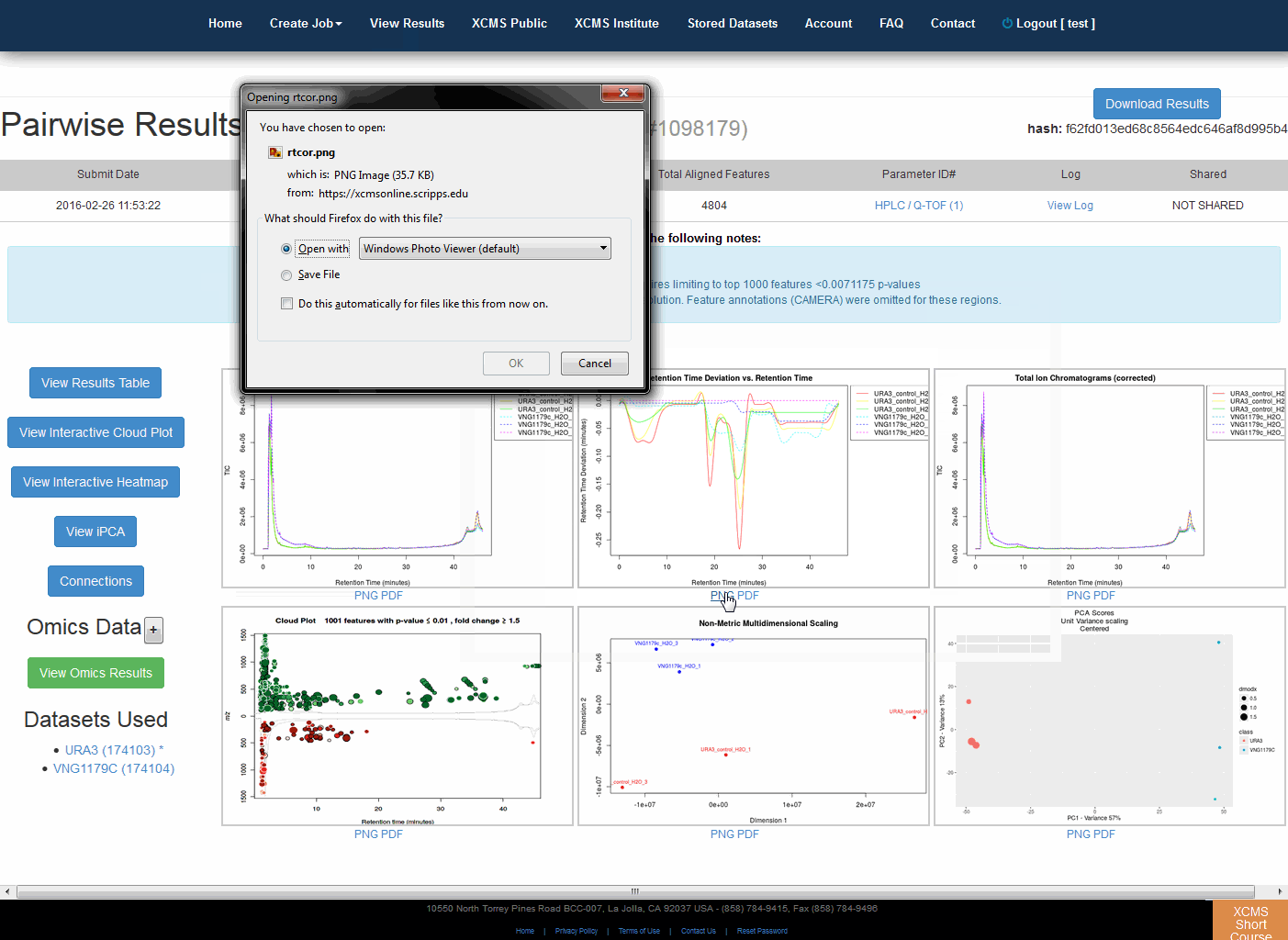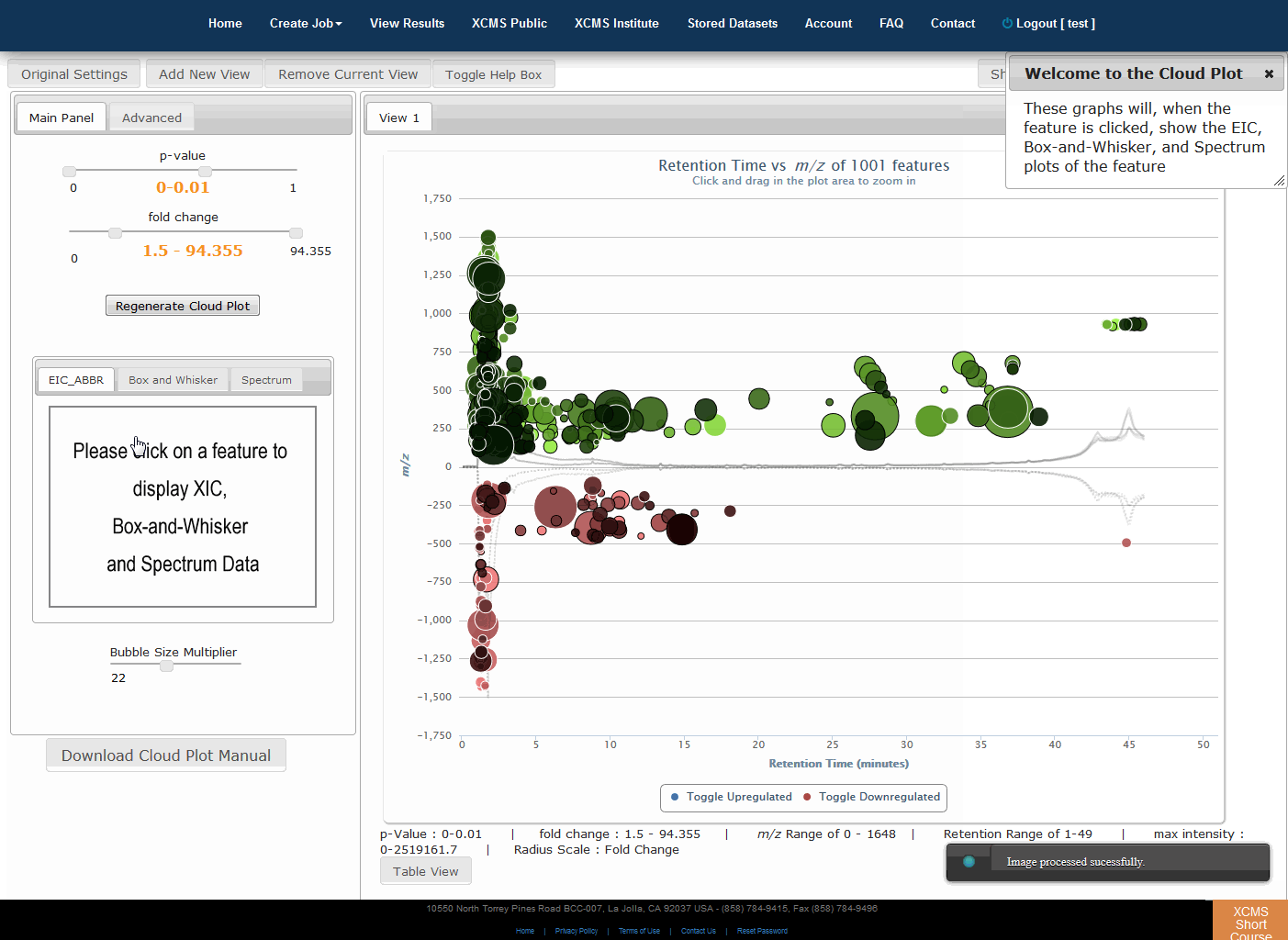
Pairwise Jobs will have an initial results screen that provides a summary with the following graphs:
Total Ion Chromatograms (TIC) Before retention time alignment
Retention Time Deviation vs Retention Time
LTotal Ion Chromatograms (TIC) post retention time correction
Cloud plot
Multidimensional Scaling (MDS)
Principal Component Analysis (PCA)
Optionally a MS/MS scan location plot if MS/MS data was included in file upload
You can also download the full results as a zip file using the link at the top of the page. This file will not include any online annotations you input in the feature result table.
成对作业将有一个初始结果屏幕,其中提供包含以下图表的摘要:
保持时间对准前的总离子流色谱图(TIC)
保留时间的偏差与保留时间的偏差
保留时间校正后的总离子流色谱图(TIC)
云图绘图多维缩放法(MDS)
主成分分析(PCA)
如果文件上传中包含MS/MS数据,则可选择使用MS/MS扫描位置打印
您也可以使用页面顶部的链接下载完整的结果作为zip文件。此文件将不包括您在功能结果表中输入的任何联机注释。
Multigroup Jobs will have an initial results screen that provides the same plots as the above pairwise job.
Single Jobs will be dependent on how many files are present in the job. For example if many files are in the dataset then a retention time plot will be shown along with the corrected TIC. With multiple files a PCA and MDS plot will be shown. However, with single jobs where only one file is present then only a TIC will be shown.
metaXCMS Jobs will have an initial results screen that provides a Venn diagram of the results along with a link to the View Results Table on left-hand size of page. There are no interactive cloud, interactive heatmap or interactive PCA plots.
Multigroup Jobs将有一个初始结果屏幕,提供与上述成对作业相同的情节。
Single Jobs将取决于作业中存在的文件数量。例如,如果数据集中有许多文件,那么保留时间图将与修正后的TIC一起显示。有多个文件,一个PCA和MDS图将显示。然而,对于只有一个文件的单个作业,则只显示一个TIC。
metaXCMS Jobs将有一个初始结果屏幕,提供结果的维恩图以及页面左侧大小的查看结果表的链接。没有交互式云,交互式热图或交互式PCA图。
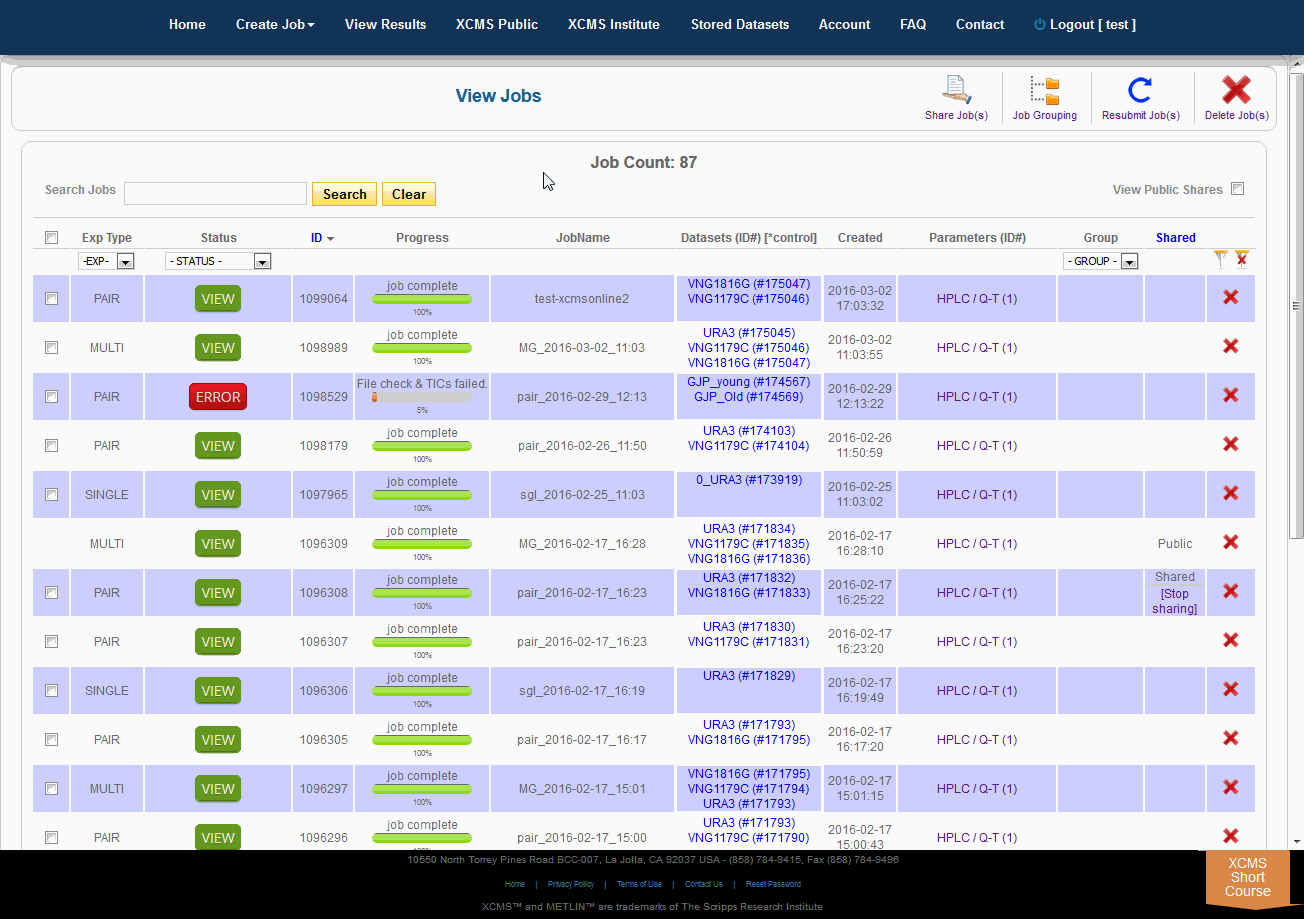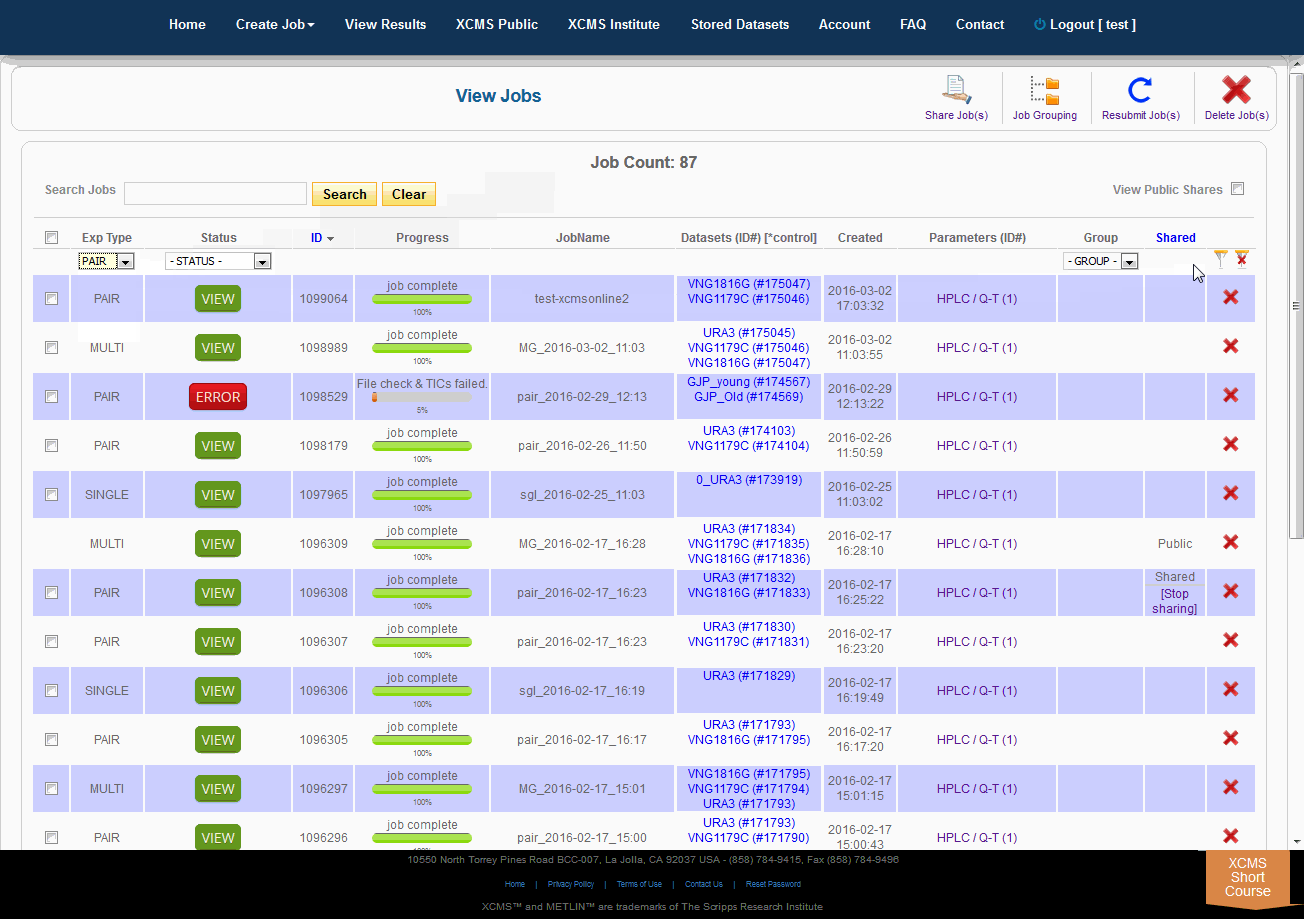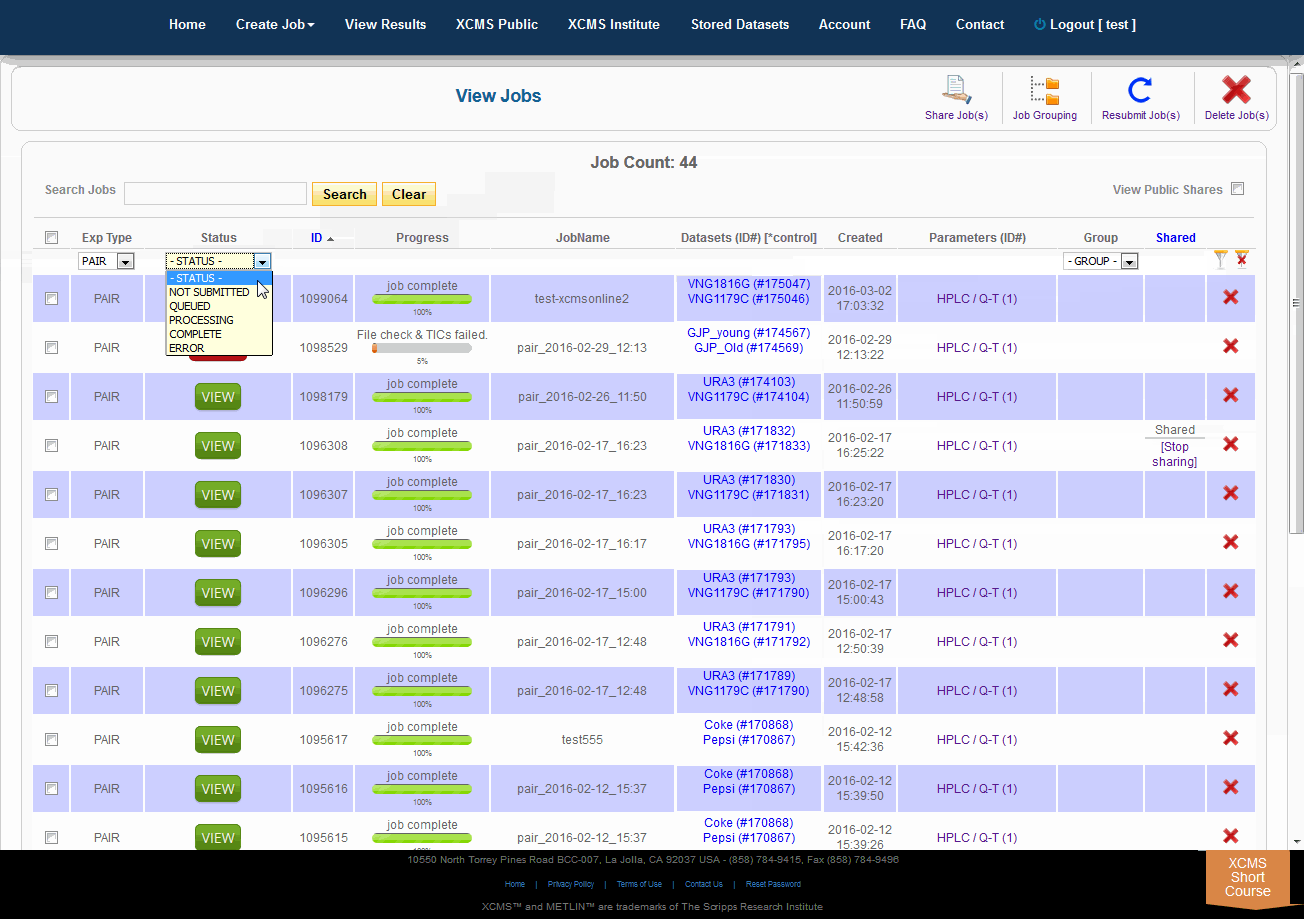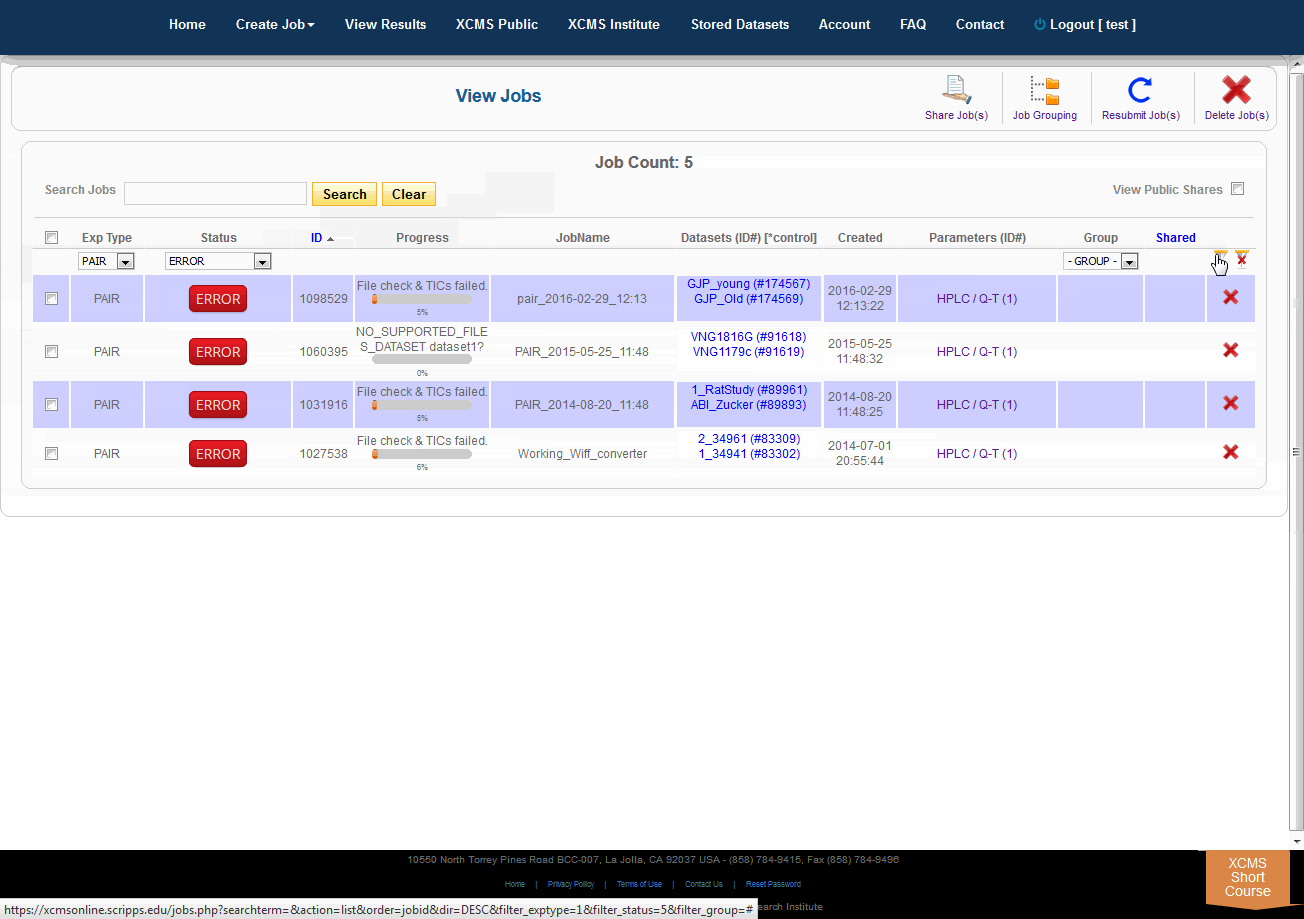
Over time you may have a large number of results on this table. Above the table is a search box that searches the columns "ID" and "JobName". Clicking the "ID" header will sort your jobs by ascending or descending order. The XCMS system also has a quick-filter mechanism to help you find data. There is a drop-down menu above the columns Exp Type, Status, and Group. Choose the values you want then click the filter button on the right side of the table. You can clear the filters with the button next to it.
随着时间的推移,您可能会在这个表上获得大量结果。表的上方是一个搜索框,用于搜索“ID”和“JobName”列。点击“ID”标题将按升序或降序排列您的工作。XCMS系统还有一个快速筛选机制来帮助您查找数据。在“Exp类型”、“状态”和“组”列上面有一个下拉菜单。选择您想要的值,然后单击表格右侧的筛选按钮。您可以使用它旁边的按钮清除过滤器。
The view results page allows you to delete a job by clicking the button in the far right column. The underlying dataset will not be deleted with a single click, as you may wish to create another job with different parameters and compare results.
查看结果页面允许您通过单击最右列中的按钮来删除作业。仅单击鼠标就不会删除底层数据集,因为您可能希望创建另一个具有不同参数的作业并比较结果。
3.1 查看结果表
View Results Table
The View Results Table is different than the View Results page and can be found by clicking the “VIEW” button of a job. On the left side is a button named "View Results Table." If this is the first time viewing the current job, the data will be loaded into the database for viewing and annotating which may cause a brief delay. Subsequent viewing will be faster. As you navigate through each feature (you can use arrow keys), details about each feature is displayed to the right of the table (m/z, rete ntion time, EIC, box-and-whisker plot and putative METLIN ID’s).Multigroup jobs will have an additional tab in the Mass-Spectrum/Box-and-Whisker plot section for post-hoc calculations between datasets.
“查看结果”表不同于“查看结果”页面,可以通过单击作业的“查看”按钮找到。在左边是一个名为“查看结果表”的按钮。如果这是第一次查看当前作业,则数据将被加载到数据库中以进行查看和注释,这可能会导致短暂的延迟。随后的查看速度将会更快。浏览每个功能时(可以使用箭头键),每个功能的详细信息显示在表右侧(m/z、保留时间、EIC、框图和假定的METLINID)。
Searching/filtering is available by clicking the magnifying glass icon lower-left of the table.
You can hide or display specific columns by clicking the icon in the lower-left.
Clicking any column header will sort the table in ascending or descending order by that field.
Column order and width can be configured by dragging and dropping column headers.
Double clicking a row or pressing “Enter” will open the notes dialog box. This information is saved in notes column.
多组作业将在“质谱/框和耳语”图部分有一个附加选项卡,用于数据集之间的事后计算。
可通过单击表格左下方的放大镜图标进行搜索/过滤。
您可以通过单击左下方的图标来隐藏或显示特定的列。
单击任何列标题将按该字段按升序或降序对表进行排序。
可以通过拖放列头来配置列的顺序和宽度。
双击一行或按Enter键将打开笔记对话框。此信息将保存在注释列中。
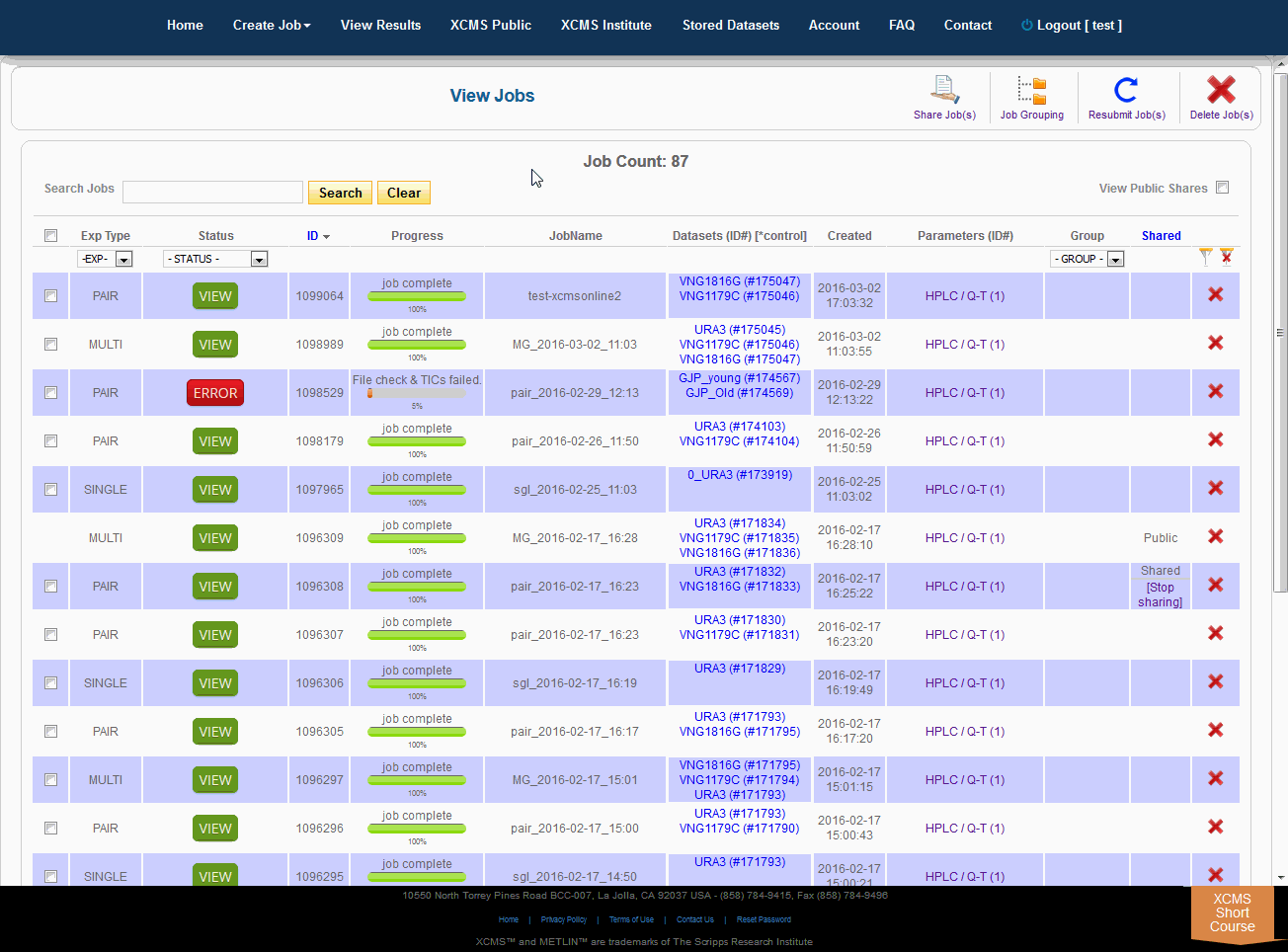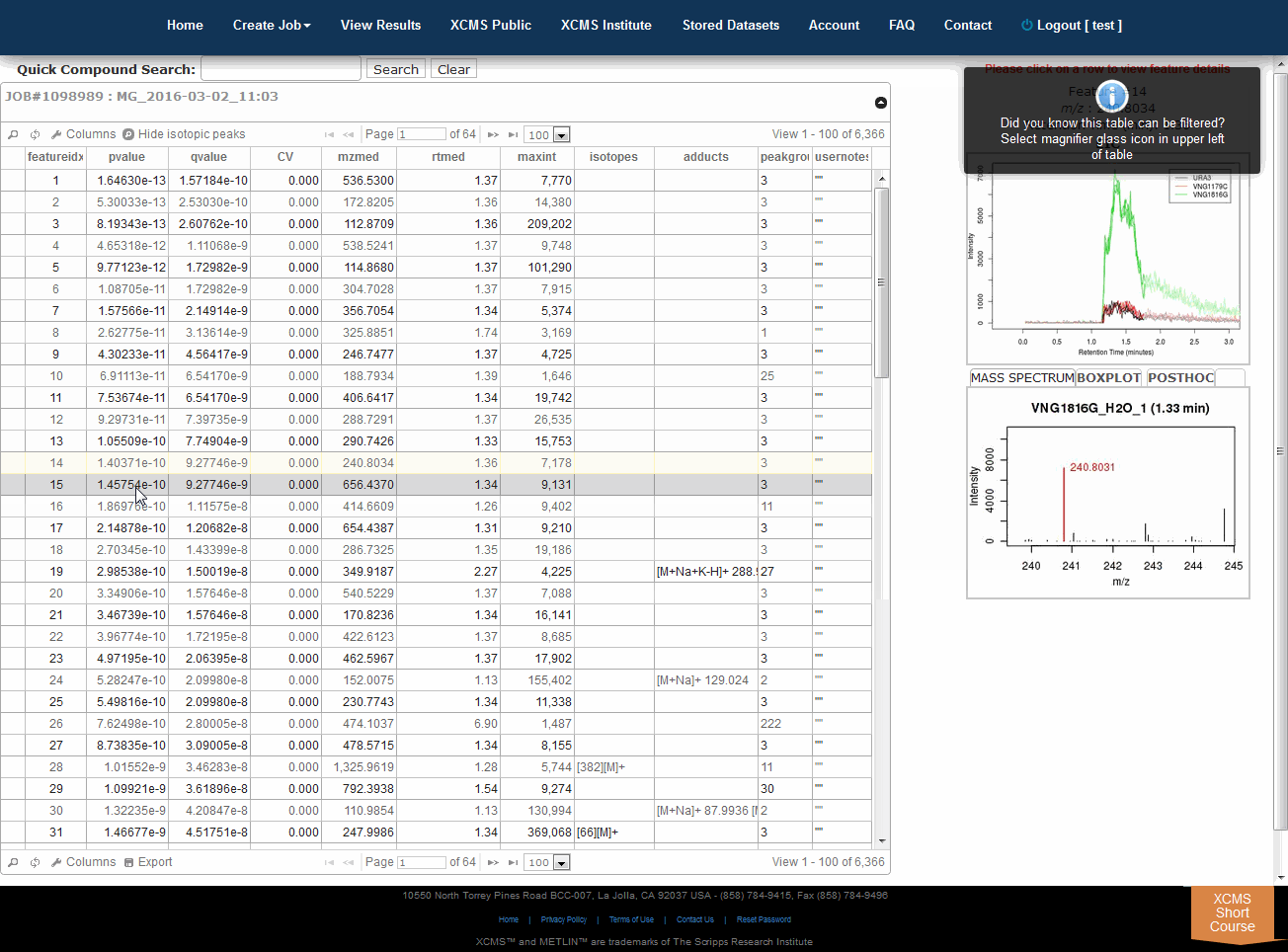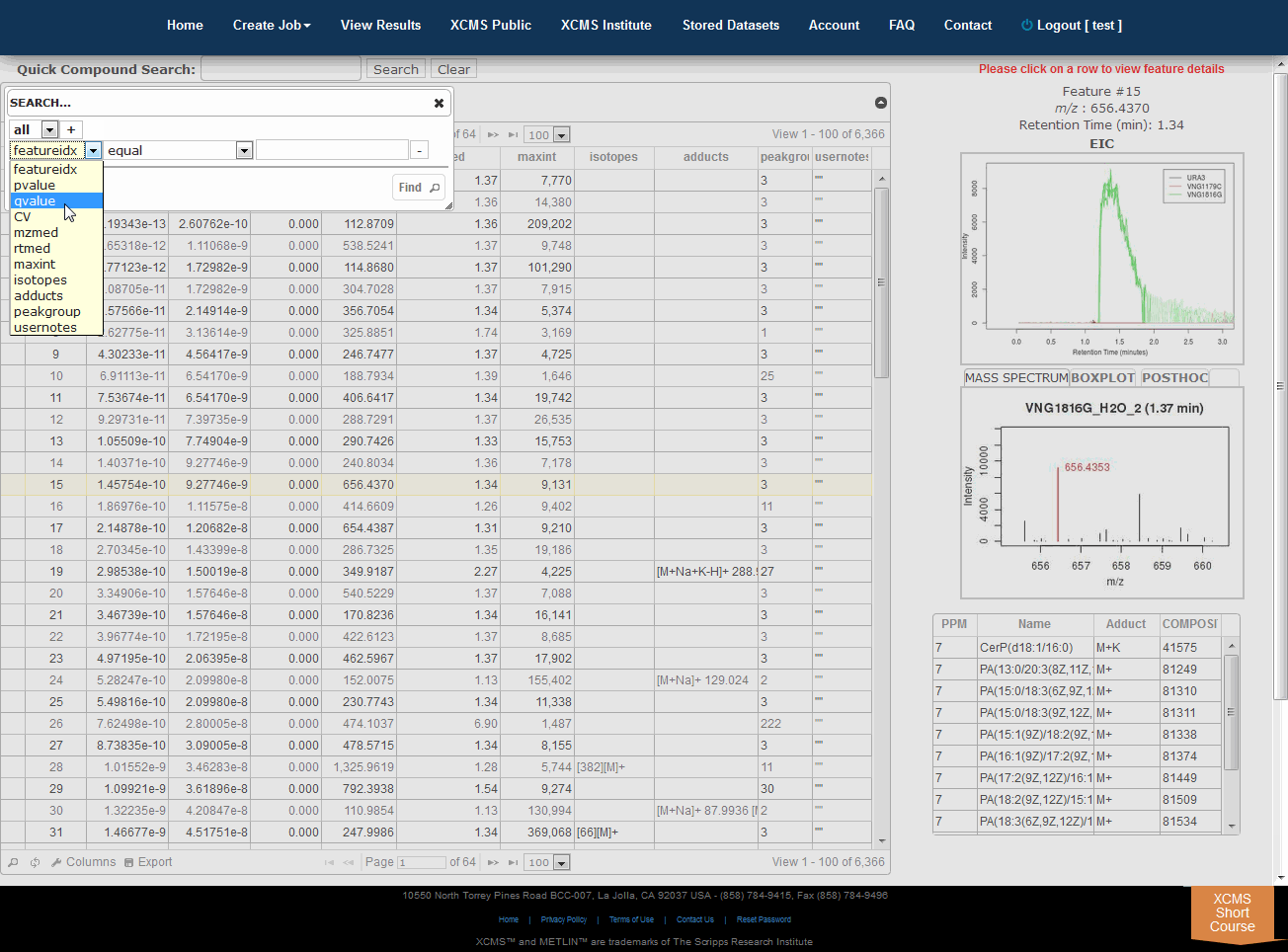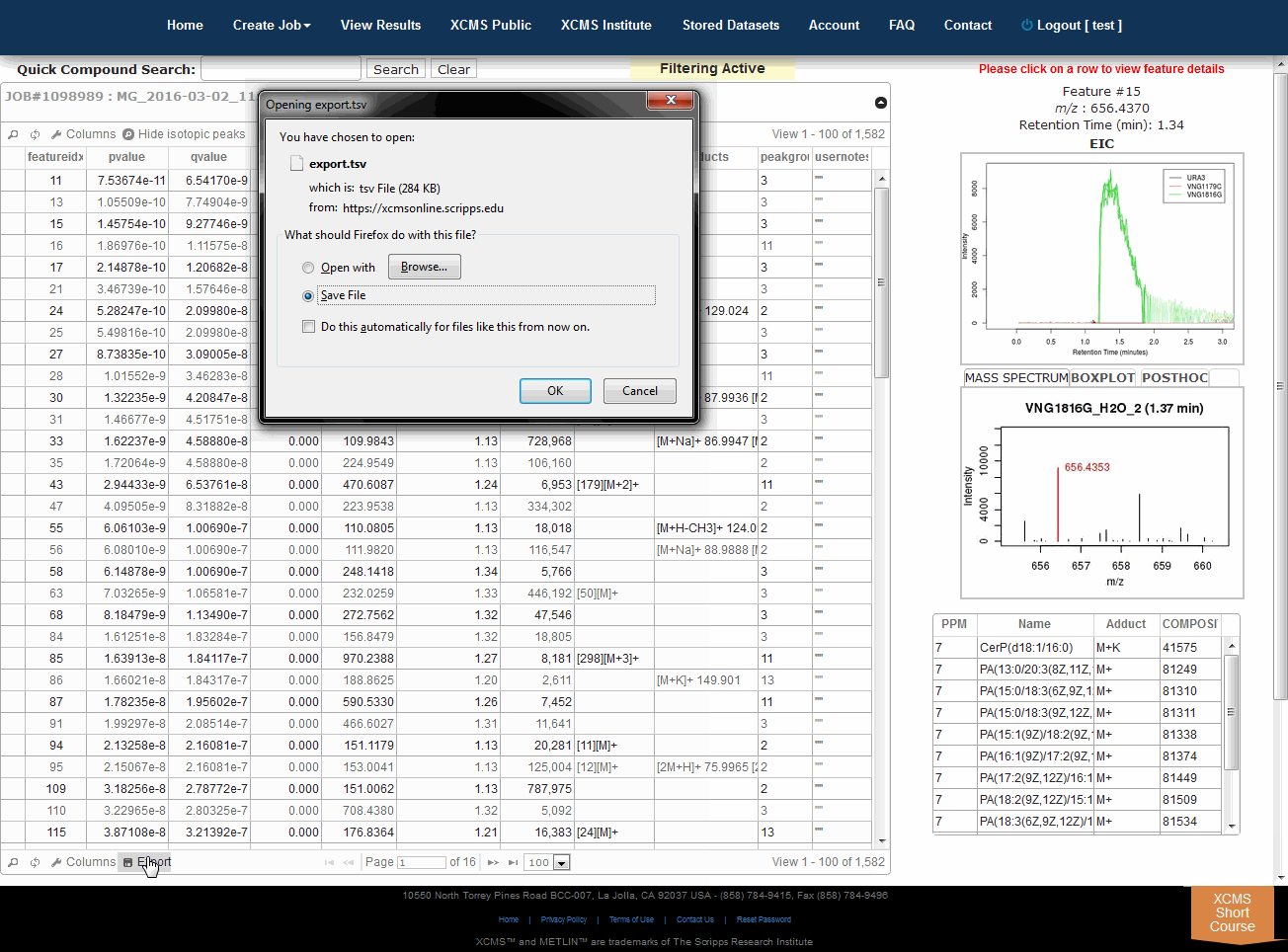
3.2 交互式云图
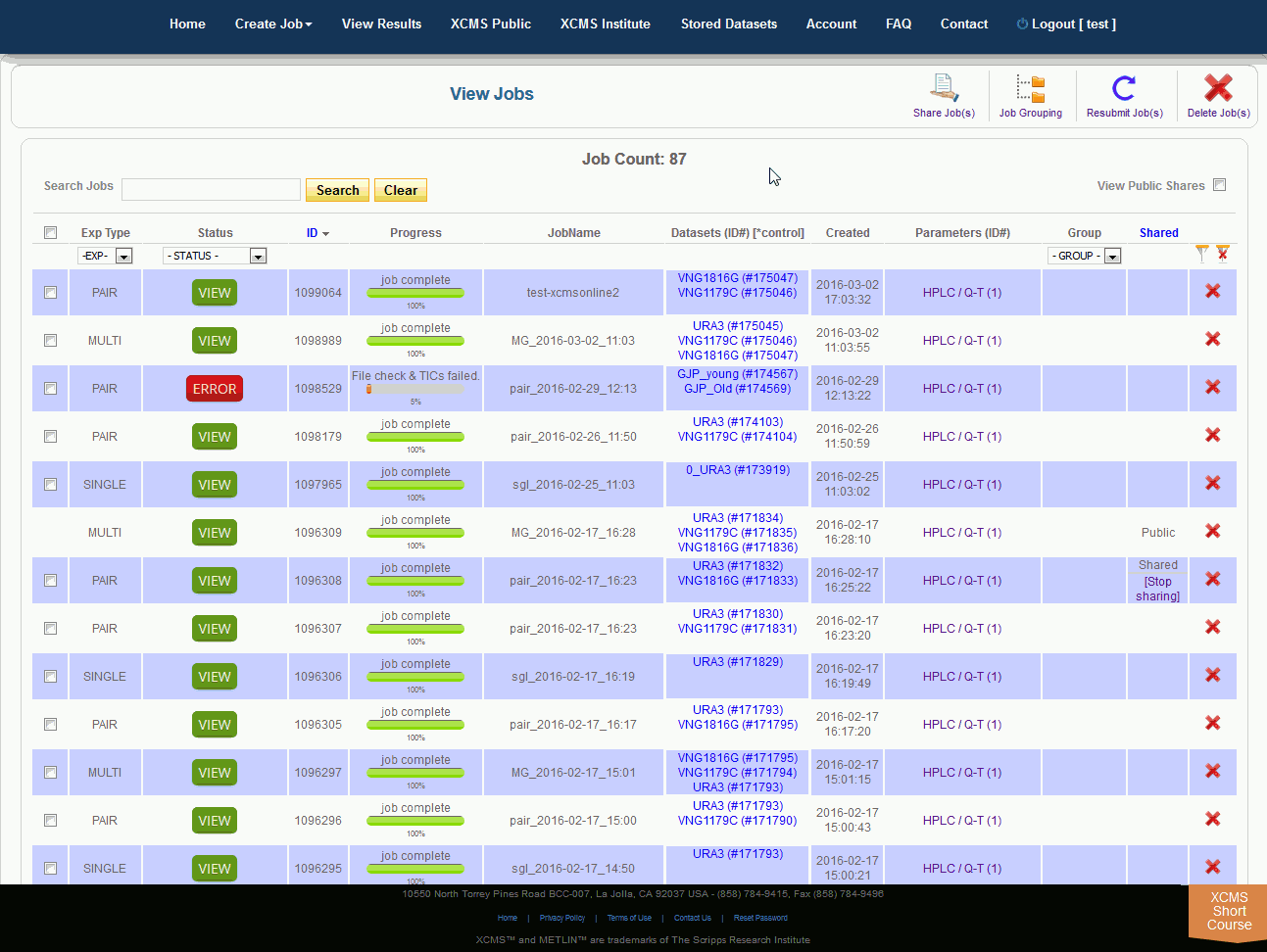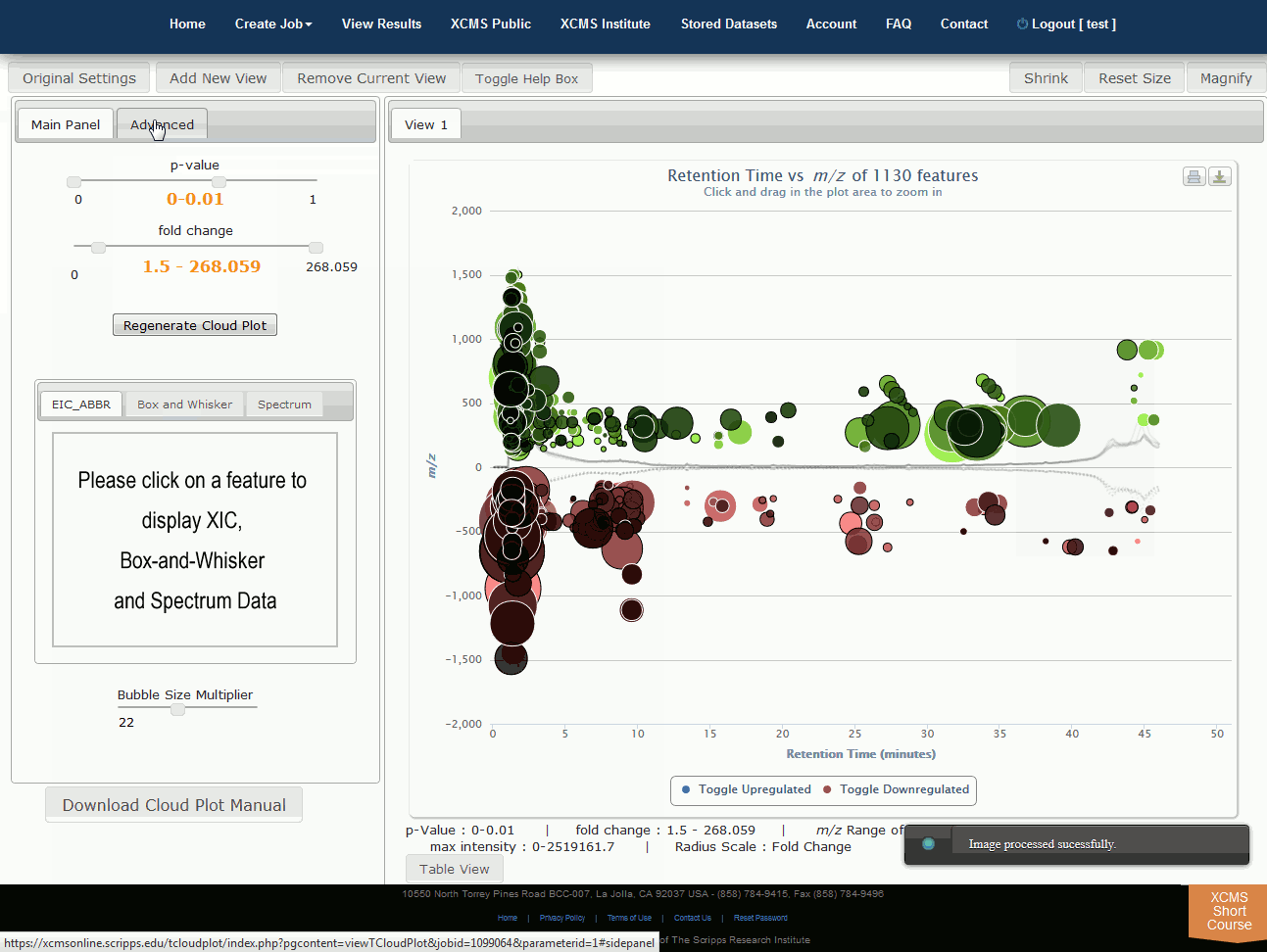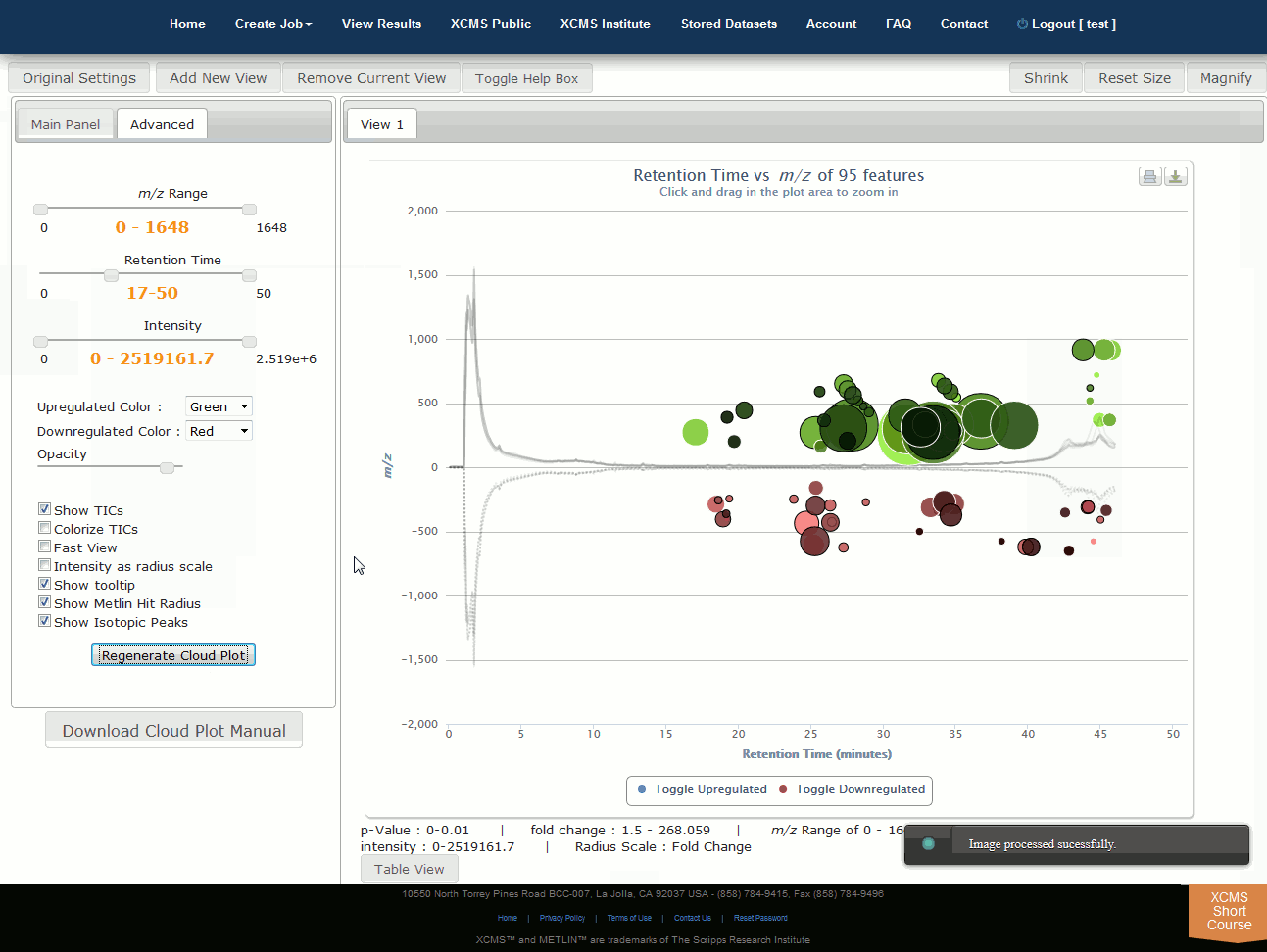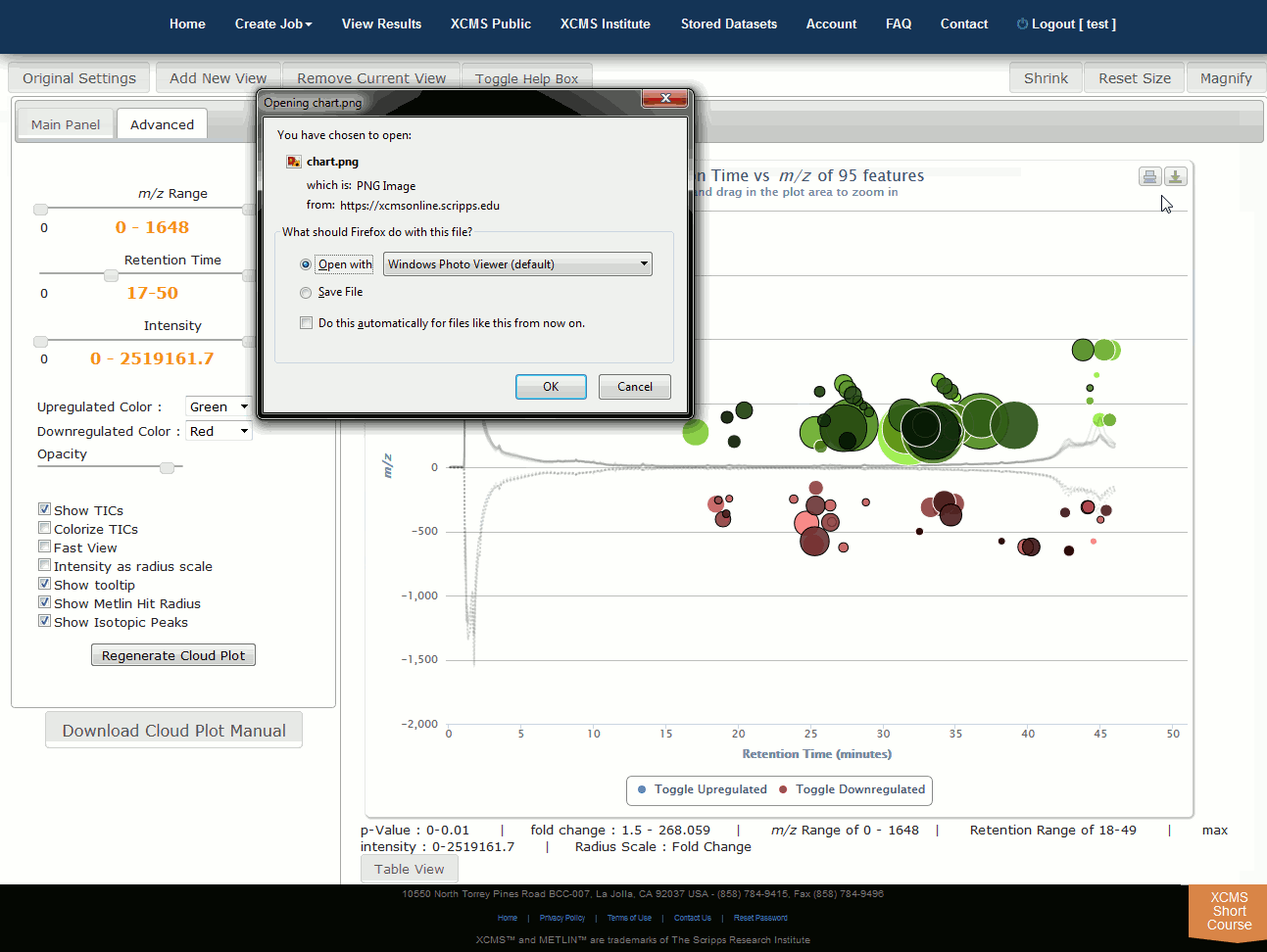
Interactive Cloud Plot
Click “View Interactive Cloud Plot” button. The interactive cloud plot will display all features as “bubbles” in the right pane while details can be found in the left pane. You can adjust parameters on the left then click “Regenerate Cloud Plot” to see the results. You can see this information in a table by clicking the "Table" button under the plot.
You can print the cloud plot or export it as in image for use in publications by clicking the icons in the upper-right corner of the plot. If you use any of the features or images in XCMS Online as part of a publication, we request you cite the appropriate source publication upon which the technology is based. Please see the Citation Reference section in Appendix II for publication references.
NOTE: Detailed information for the cloud plot feature is available in the "Cloud Plot Manual" button in lower-left of the screen.
点击“查看交互式云图”按钮。交互式云图将在右窗格中显示所有特征为“气泡”,同时也可以在左窗格中找到详细信息。可以调整左侧的参数,然后点击“重新生成云图”,查看结果。您可以通过点击地块下的“表格”按钮来在表格中看到此信息。
可以通过单击打印右上角的图标来打印云打印,供出版物中使用。如果您使用XCMSOnline中的任何特性或图像作为出版物的一部分,我们要求您引用该技术所基于的适当的源出版物。出版物参考请见附录二的引用参考部分。
注:云图功能的详细信息,见屏幕左下角的“云图手册”按钮。
3.3 互动 PCA
Interactive PCA
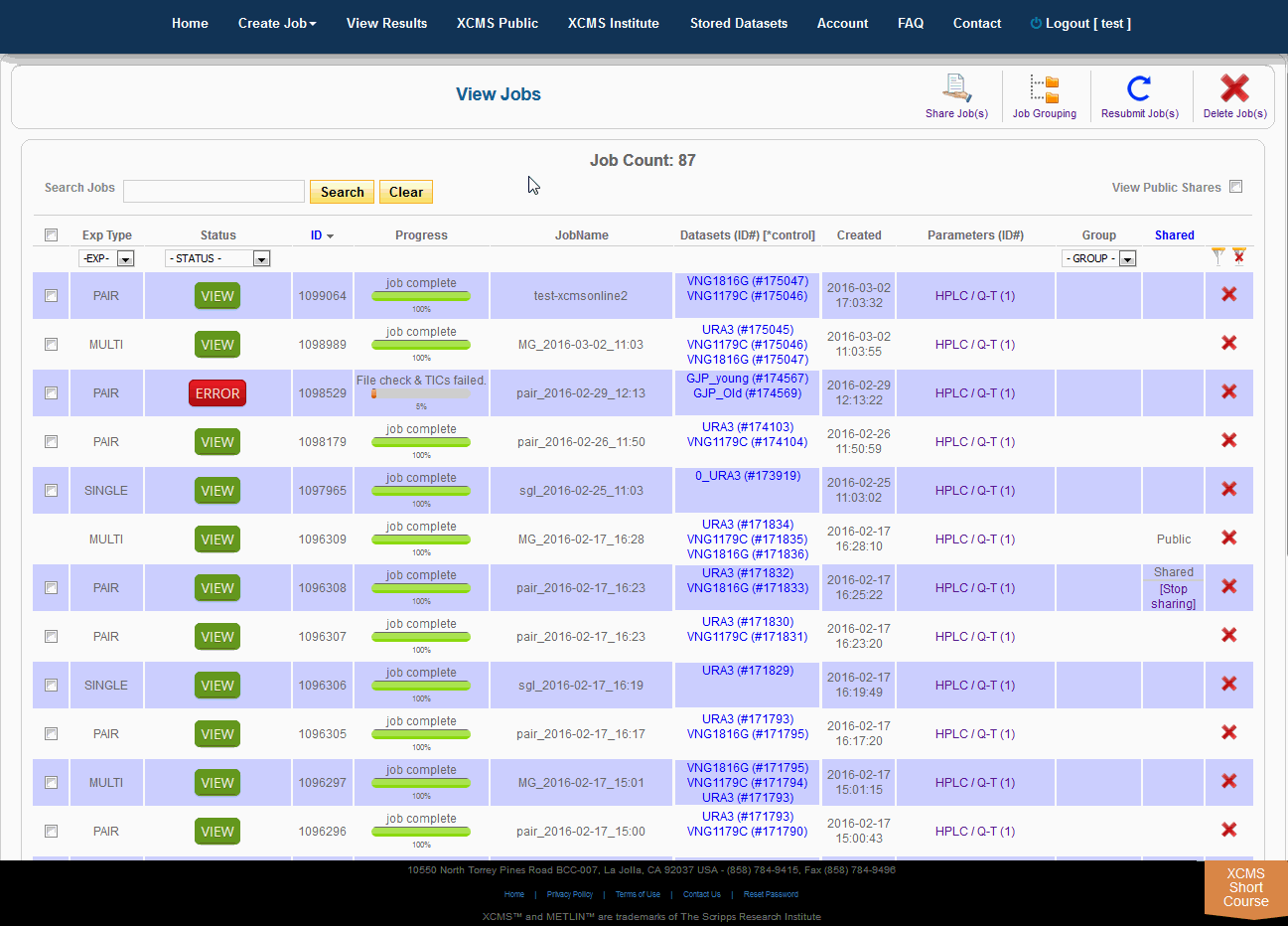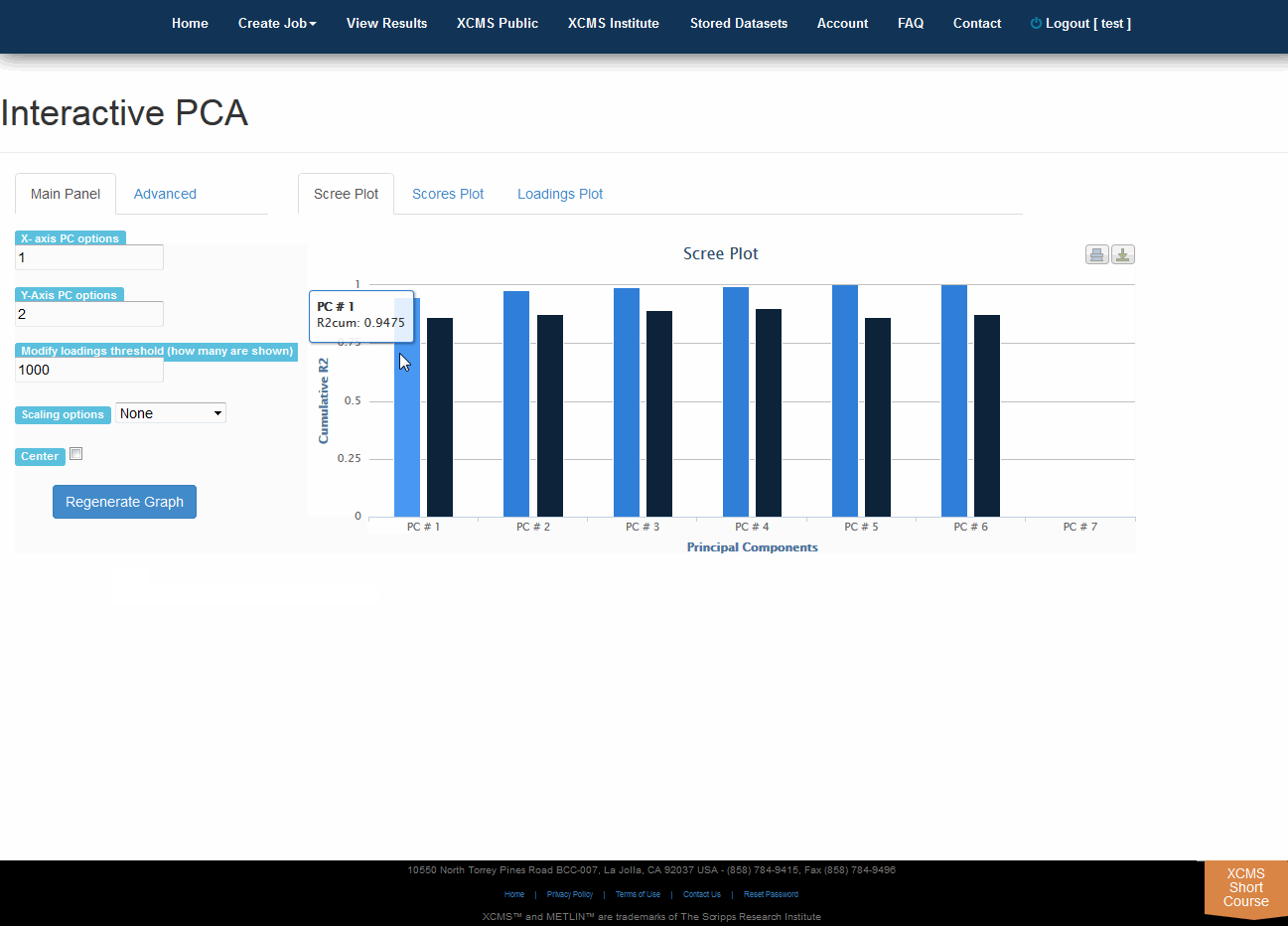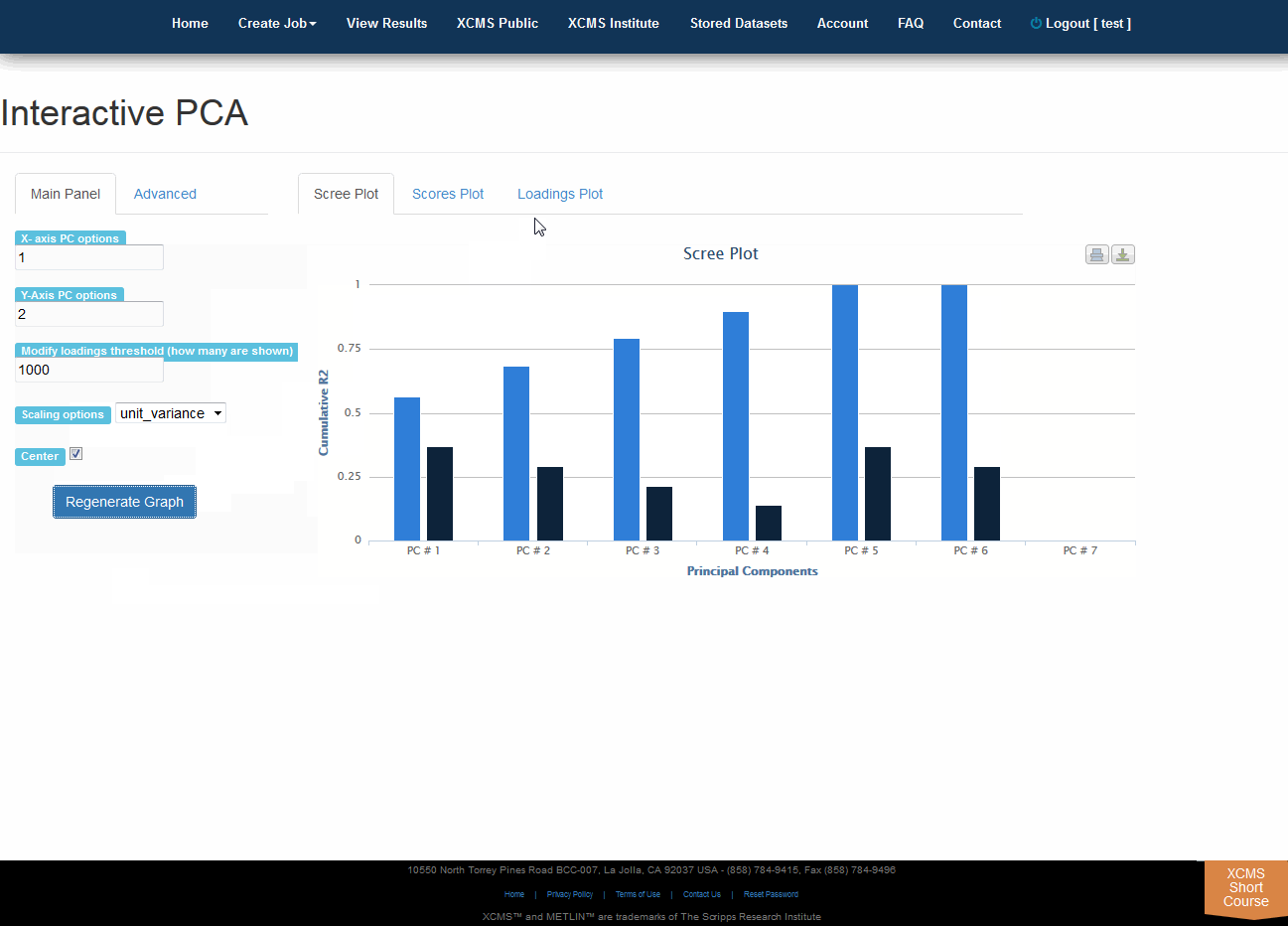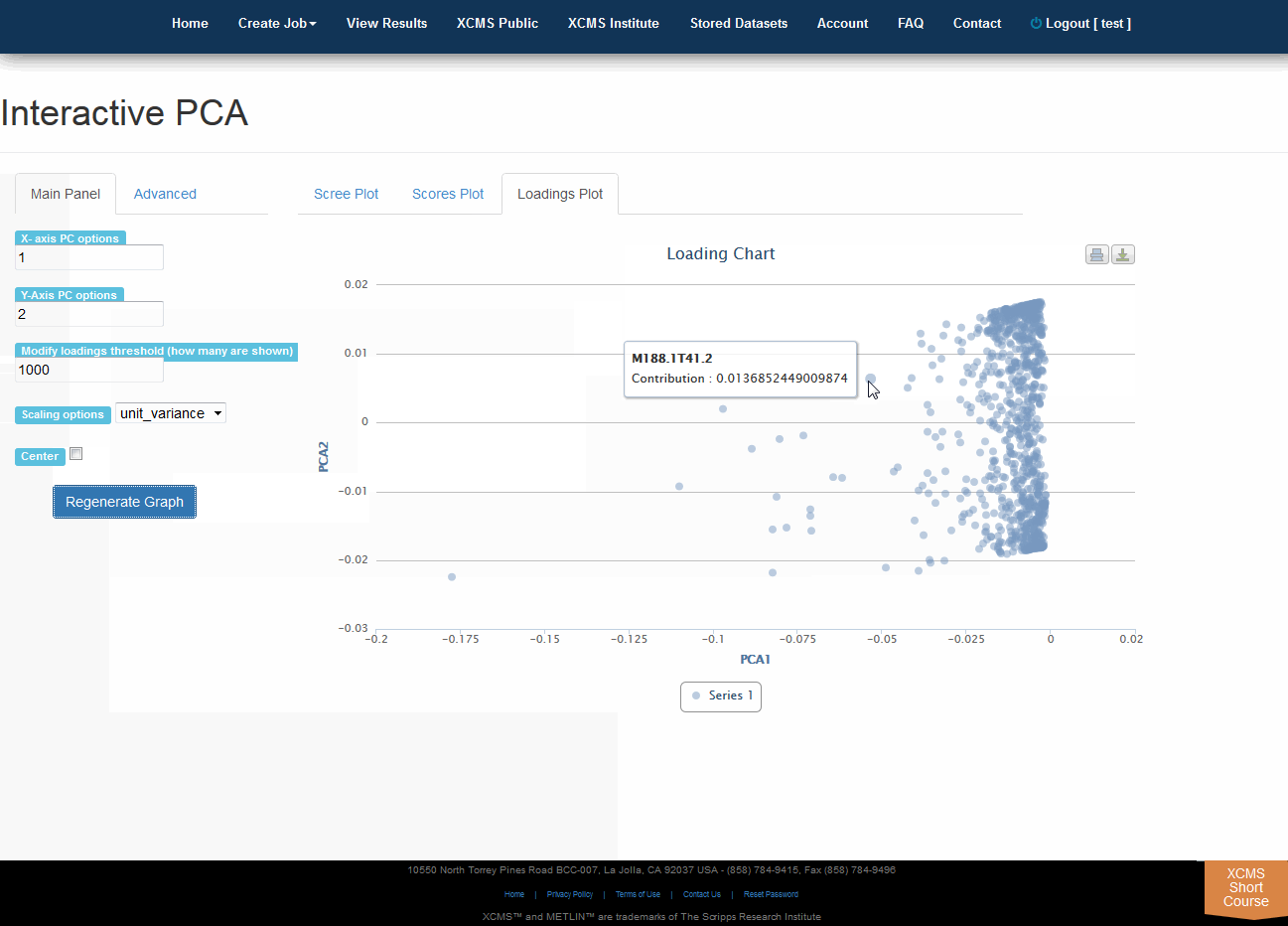
Click “View Interactive PCA” button to View Interactive Principal Component Analysis. The iPCA feature allows the user to modify the axes, loadings and export the resulting scree, scores or loadings plots.
点击“查看交互式PCA”按钮,查看交互式主体组件分析。iPCA功能允许用户修改轴、加载和导出生成的屏幕、分数或加载图。
3.4 交互式热图
Interactive Heatmap
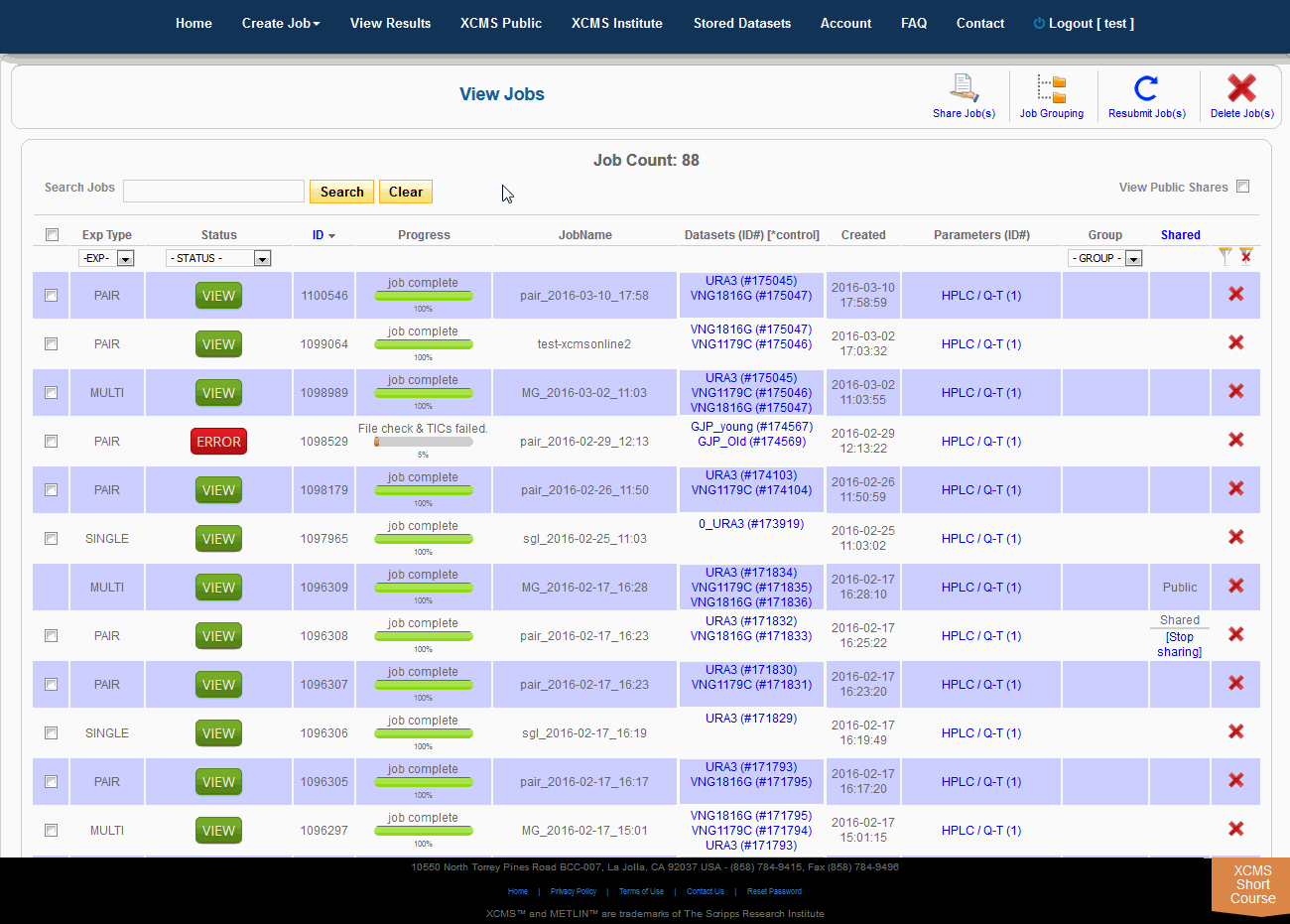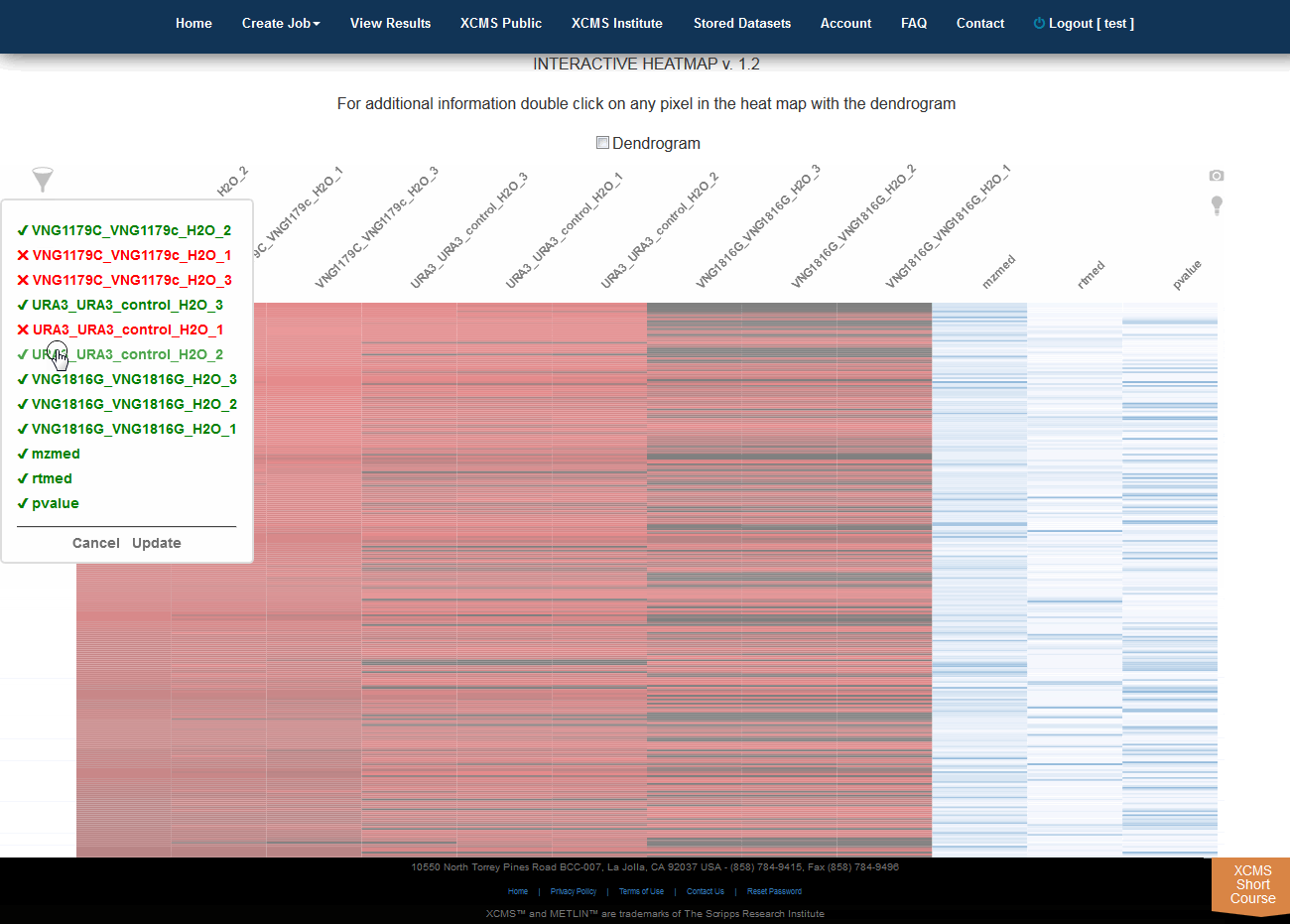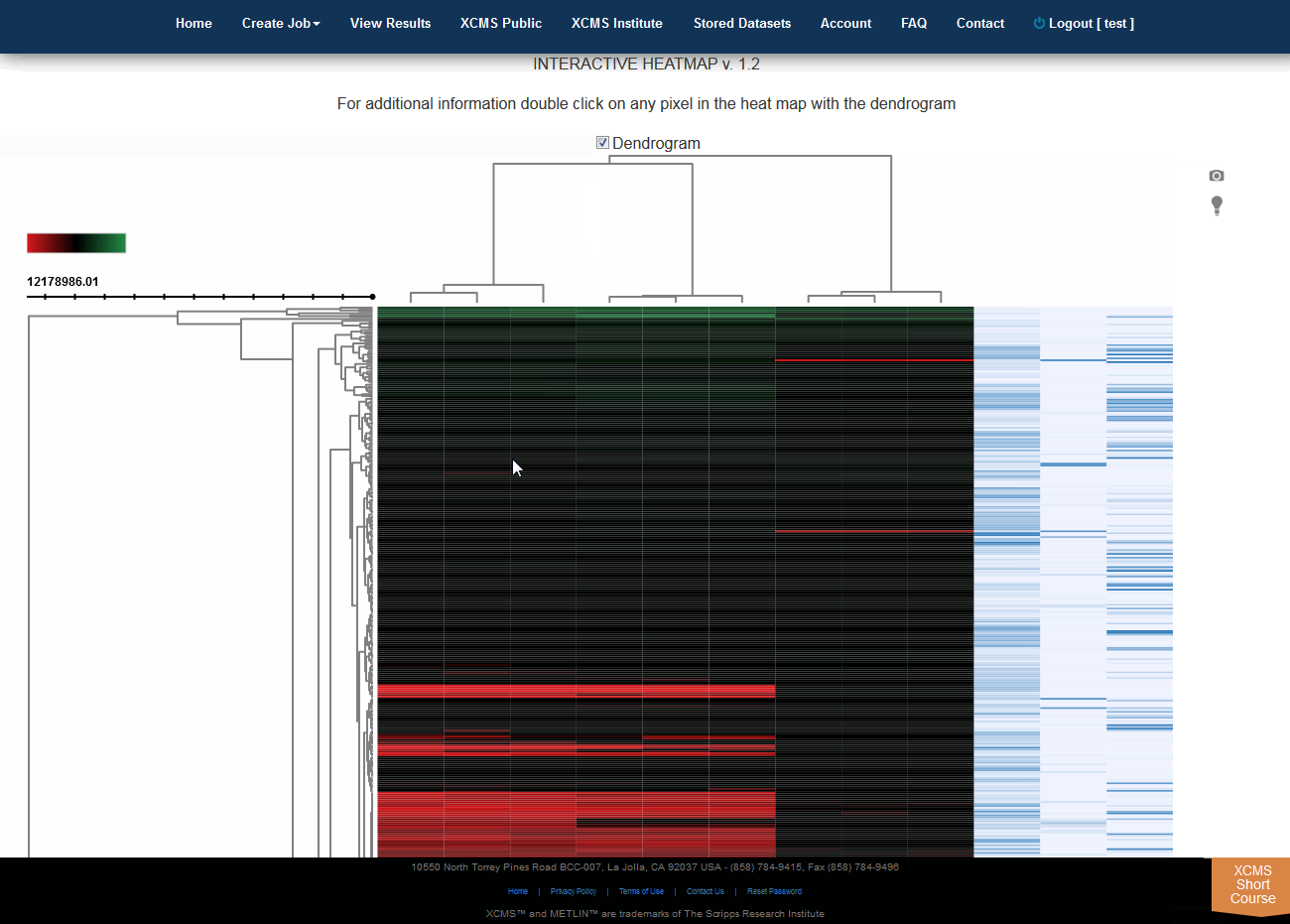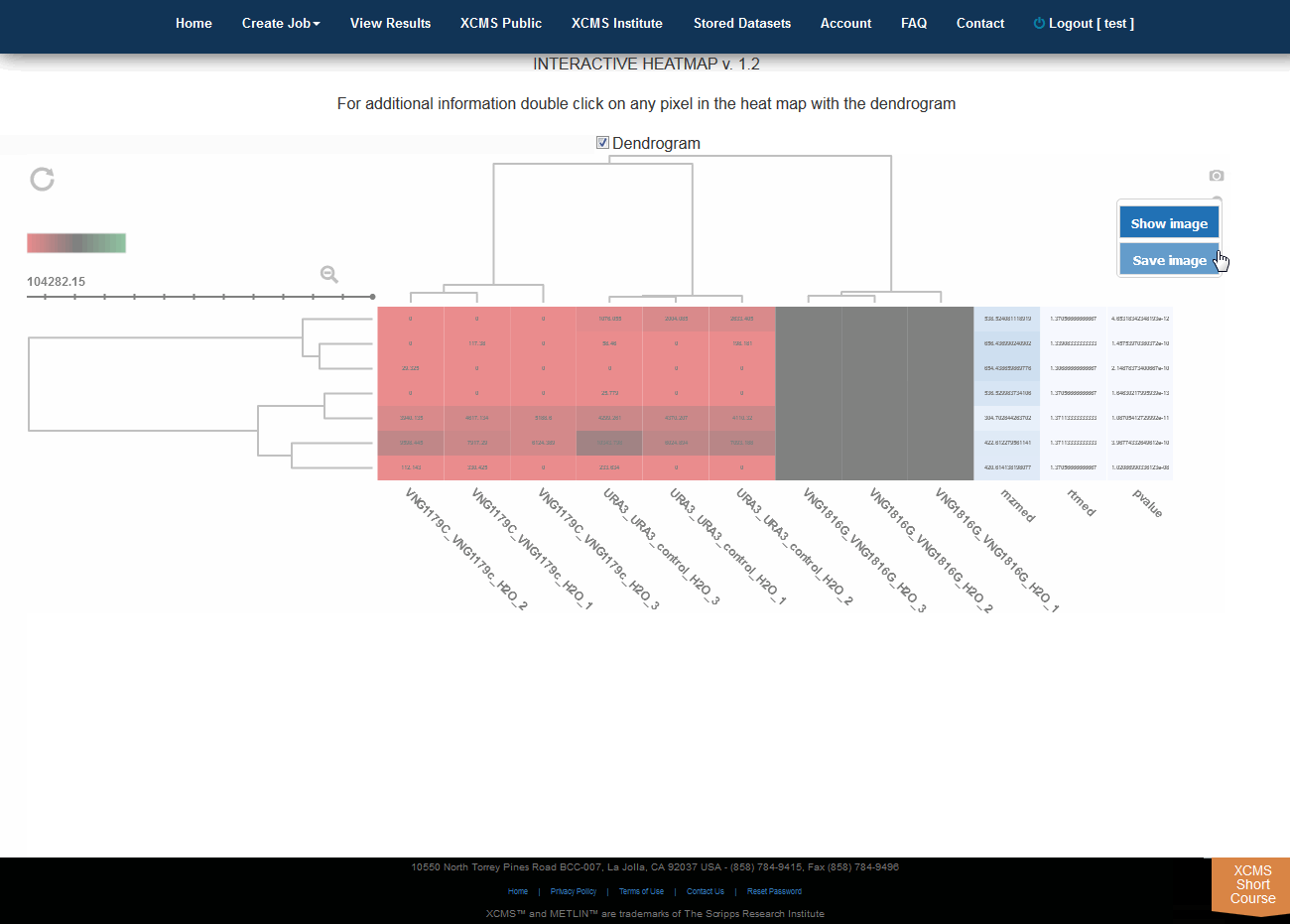
Click “View Interactive Heatmap” button to view Interactive Heatmap. The interactive heatmap displays the data in a format that can be analyzed visually based on color but may also be sorted/zoomed to display additional detail about specific features. Detail is displayed when the mouse rolls over the box. Clicking the box will open a separate window where feature number, meta information, graphs (box-whisker, EIC, mass spectrum) and possible METLIN hits are displayed. You may have to permit popup windows in your browser settings for this feature to work correctly.
点击“View Interactive Heatmap”按钮,查看交互式热图。交互式热图以一种格式显示数据,该格式可以根据颜色进行可视化分析,但也可以进行排序/缩放,以显示有关特定特征的其他细节。鼠标滚动时显示详细信息。单击该方框将打开一个单独的窗口,其中显示特征编号、元信息、图形(方框搅拌器、EIC、质谱)和可能的METLIN点击量。您可能必须允许浏览器设置中的弹出窗口,这样此功能才能正常工作。
3.5 下载数据
Downloading Data
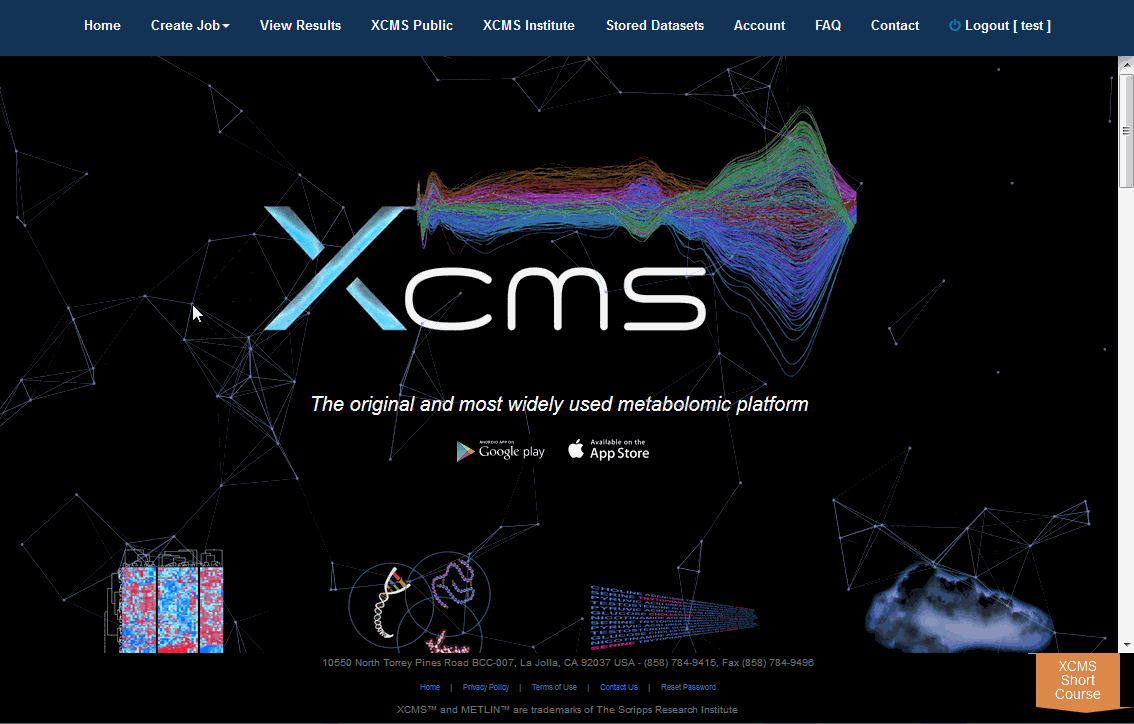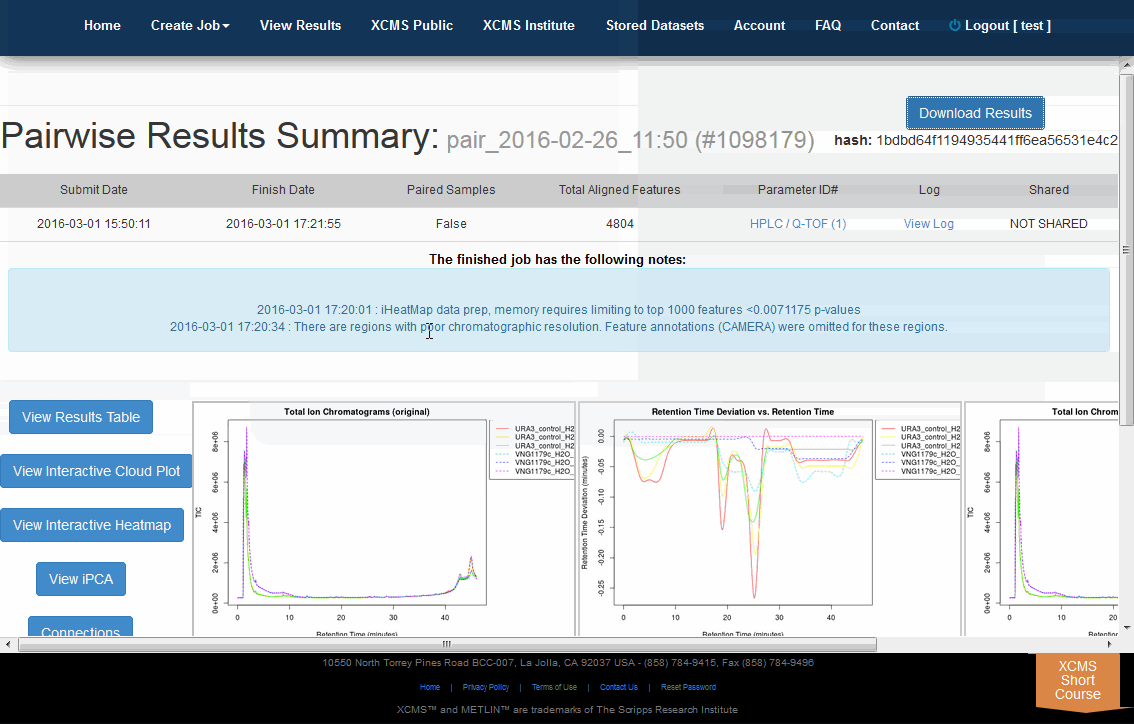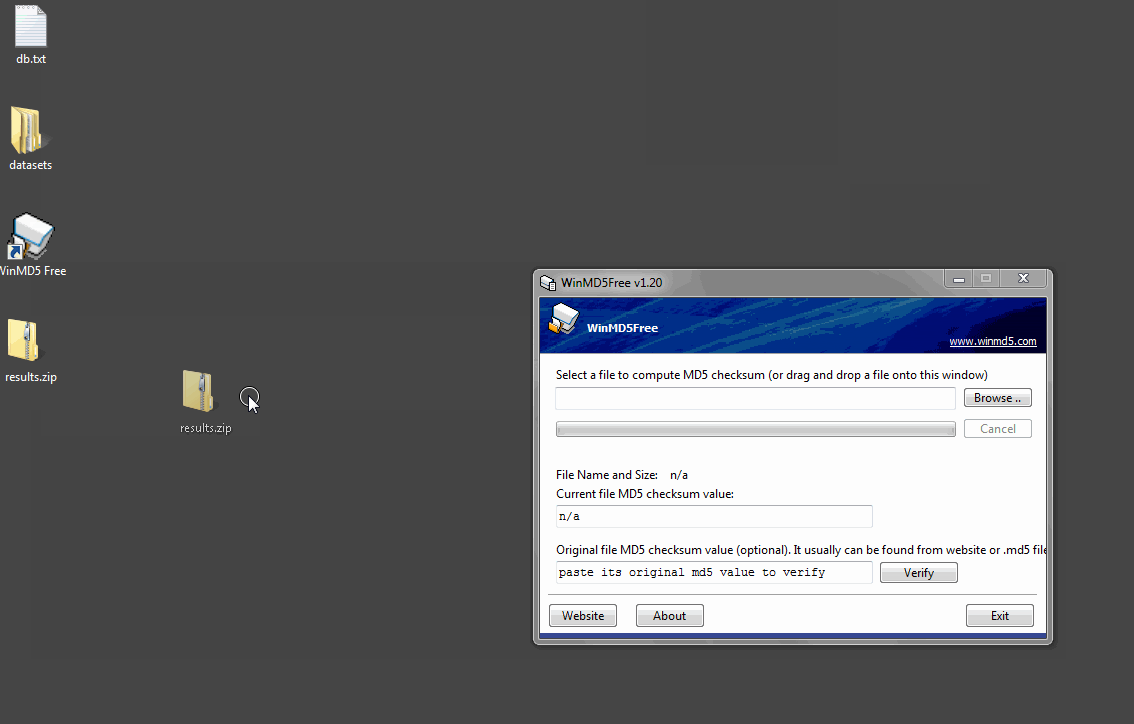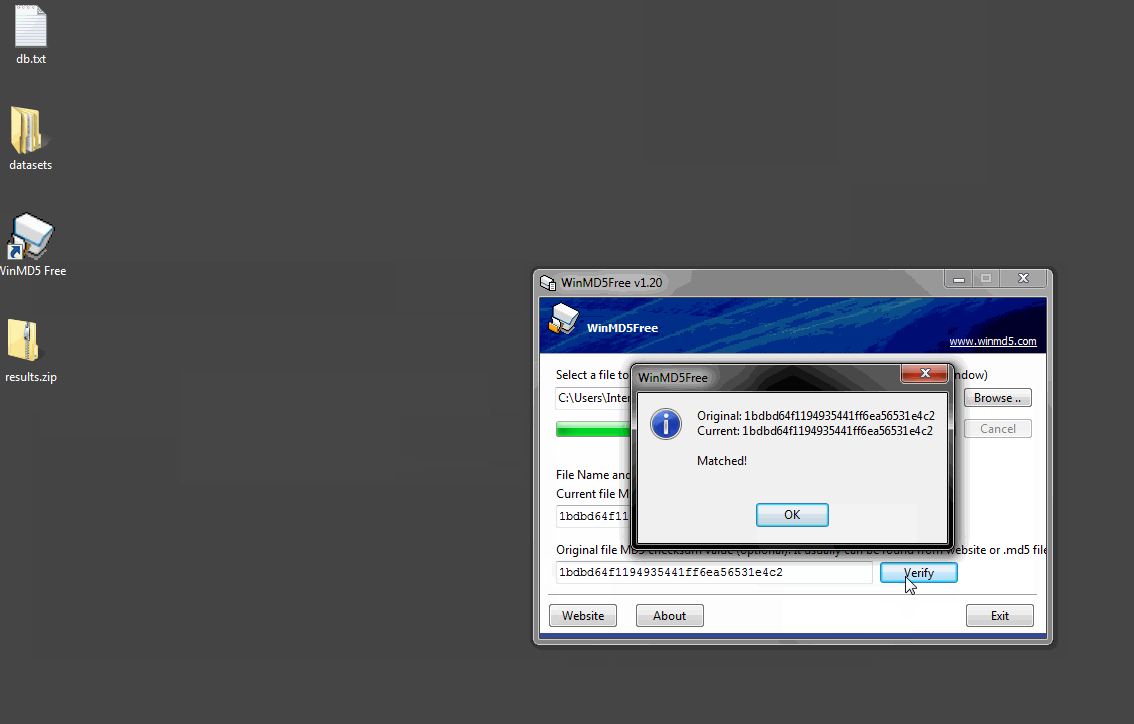
Click "View Results" from the top navigation menu then "view" your desired experiment. Click "Download Results" from the upper right corner. This will begin the download of a file named results.zip. Below the button is the word hash: followed by a long string of characters. This is a MD5 checksum value, a cryptographically generated number based on the input file. Think of it as a unique signature for your file. We use MD5 hashing to verify data integrity during transfer. If you want to verify the data has not been corrupted during transfer use a MD5 hash tool to compare the downloaded file with this number.
For Windows users we recommend a tool named WinMD5 Free. (http://winmd5.com/) Download the file named windmd5free.zip and extract the executable named WinMD5.exe. There is no installer, you simply double-click the program to run it.
点击顶部导航菜单中的“查看结果”,然后“查看”您想要的实验。从右上角点击“下载结果”。这将开始下载一个名为results.zip的文件。按钮下面是单词哈希:后面是一长串字符串。这是一个MD5校验和值,一个基于输入文件的加密生成的数字。请将其视为文件的唯一签名。我们使用MD5哈希来验证传输期间的数据完整性。如果要验证传输期间数据没有损坏,请使用MD5哈希工具将下载的文件与此数字进行比较。
对于Windows用户,我们推荐一个名为WinMD5Free的工具。(http://winmd5.com/)下载名为windmd5free.zip的文件,并提取名为WinMD5.exe的可执行文件。没有安装程序,您只需双击该程序即可运行它。

Once the program is open, click the Browse button and navigate to results.zip or drag the file onto the window. WinMD5 Free will calculate and display the signature. Copy and paste the hash value from the XCMS website into the bottom field and click Verify. WinMD5 will tell you if there is a match.
程序打开后,单击“浏览”按钮并导航到results.zip或将文件拖动到窗口中。WinMD5Free将计算并显示签名。将XCMS网站中的哈希值复制并粘贴到底部字段中,然后单击“验证”。WinMD5会告诉你是否有匹配项。
4.0 账户管理
Account Management
Storage of user data is limited. This includes all uploaded datasets, custom parameters and jobs results (including annotations). To free space for additional uploads, you will likely need to delete jobs and/or datasets. To check your account storage, navigate to the Account menu. The sub navigation menu along the bottom of the main navigation has miscellaneous settings affecting your account such as "Alerts" and "Reporting." The “User Settings” allows you to modify the default theme used for displaying job results. The drop-down preview menu shows representative display.
用户数据的存储能力很有限。这包括所有上载的数据集、自定义参数和作业结果(包括注释)。要释放空间进行上载,您可能需要删除作业和/或数据集。若要检查帐户的存储空间,请导航到“帐户”菜单。沿着主导航底部的子导航菜单包含影响您的帐户的其他设置,如“警报”和“报告”。“用户设置”允许您修改用于显示作业结果的默认主题。下拉下的预览菜单显示了具有代表性的显示。
5.0 数据集管理
Dataset Management
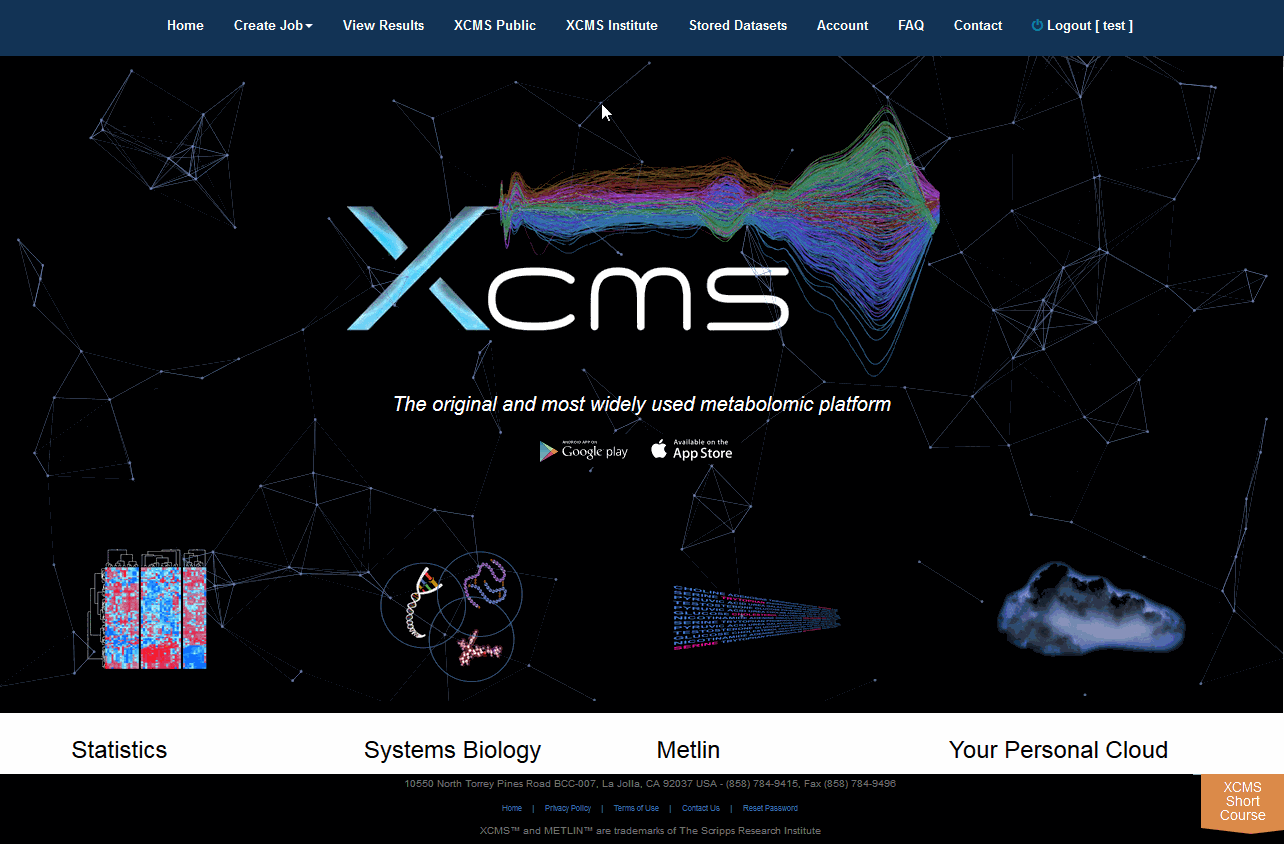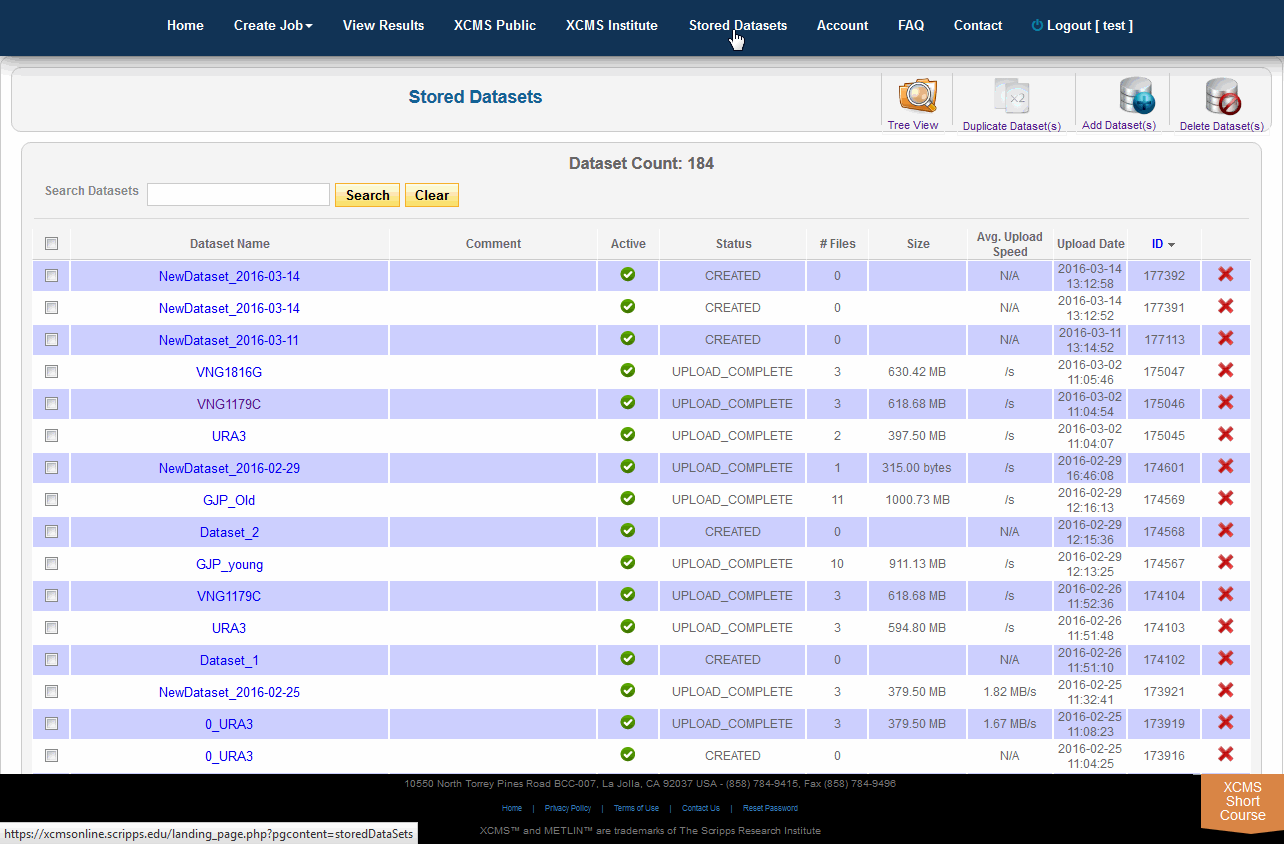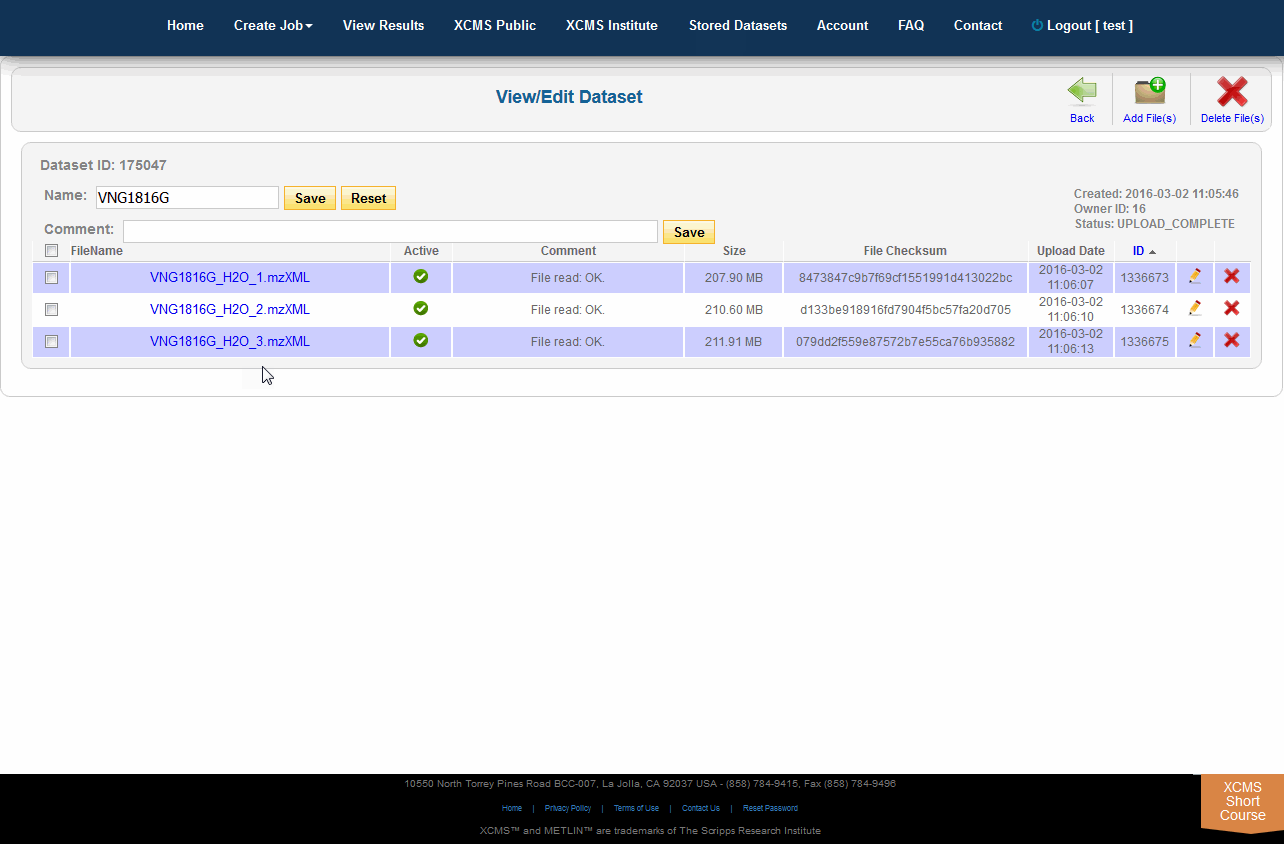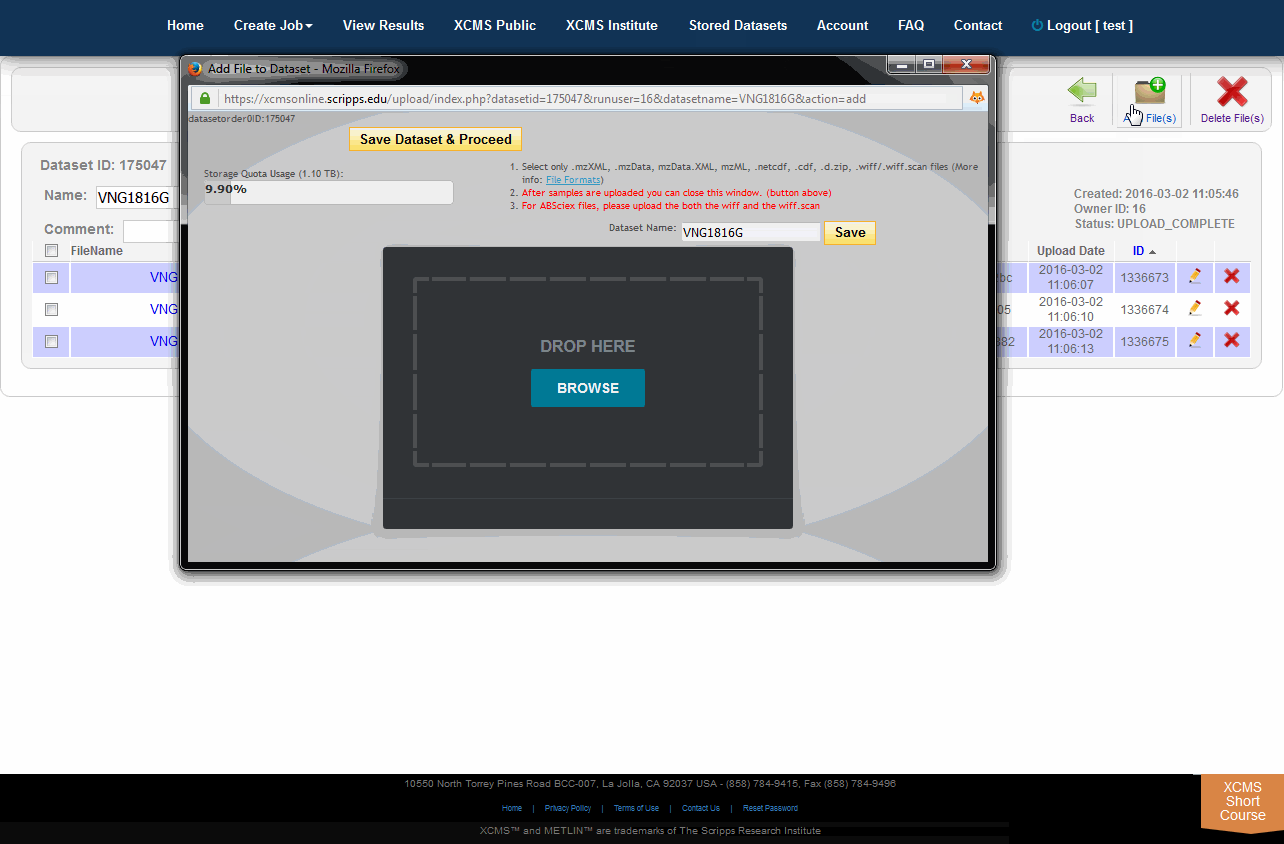
Click on “Stored Datasets” in the top navigation bar to access the stored datasets area. This page will list all datasets previously uploaded. From here you may delete the entire dataset by clicking the “X” in the far right column. A confirmation dialog box will appear for you to continue or cancel the operation. To rerun jobs based on deleted data, you will need to re-upload the data; we cannot retrieve deleted datasets.
点击顶部导航栏中的“存储数据集”,访问存储数据集区域。此页面将列出以前上载的所有数据集。从这里单击最右列的“X”删除整个数据集。将显示一个确认对话框,以便您继续或取消该操作。要根据删除的数据重新运行作业,需要重新上载数据;无法检索删除的数据集。
5.1 数据集重复
Dataset Duplication
In addition to deleting, you may duplicate datasets and add files/classes to them. For duplication simply select the datasets you wish to duplicate and select “Duplicate Dataset(s)” in the sub top navigation bar. This will initiate the process, which may take several minutes depending on the underlying hardware capacity and current usage. During the copy, the status will say “COPYING” for the new dataset. Once complete, the status will automatically change to “COPIED” and the new dataset will be available for use in new job types.
除了删除之外,您还可以复制数据集并向其添加文件/类。对于复制,只需选择您希望复制的数据集,并在子顶部导航栏中选择“复制数据集”。这将启动进程,根据底层硬件容量和当前使用情况,这可能需要几分钟。在复制过程中,新数据集的状态将显示为“COPYING”。一旦完成,状态将自动变为“copy”,新的数据集将可用于新的作业类型。
5.2 将文件添加到数据集
Adding Files to Datasets
When there is a need to modify the underlying files/classes in a dataset (i.e., corrupt upload of file or forget to include a class), the user may wish to upload the missing file. If the missing file has the same name as an existing file, the existing file will be replaced with the missing one. If it is a new file the dataset will be updated. To view the files in a specific dataset, find the dataset name on the Stored Datasets page and click the name. On the following page you will find a list of all the classes in this dataset. You can then verify/add comments, check the size and change the dataset. You can add more files to this dataset by clicking the “Add File(s)” button in the sub top navigation bar.
当需要修改数据集中的底层文件/类时(例如,文件上传失败或忘记包含类),用户可能希望上传丢失的文件。如果丢失的文件与现有文件同名,则现有文件将被替换为丢失的文件。如果它是一个新文件,数据集将被更新。要查看特定数据集中的文件,请在“存储数据集”页上找到数据集名称并单击该名称。在下面的页面中,您将找到此数据集中所有类的列表。然后,您可以验证/添加注释,检查大小和更改数据集。你可以通过点击顶部导航栏的" add File(s) "按钮向这个数据集添加更多文件。
NOTE: Adding files after jobs have already completed will not re-run them. The jobs that rely on modified dataset(s) will need to be re-run manually.
注意:在作业已完成后添加文件将不会重新运行它们。依赖修改数据集的作业需要手动重新运行。
5.3 树状图视图
Tree View
XCMS Online allows you to easily create a unique dataset from any combination of existing files using the Tree View. To do this, select "Stored Datasets" from the top navigation menu the click "Tree View" in the upper right sub-menu. All your datasets will be listed in a file explorer on the left side of the screen. You can select whole datasets by clicking the check-box to the left of each name, or select partial data by clicking the triangle to unfold the related files. Once you have selected the desired data, enter an unique name for your new dataset in the "Dataset name" field then click "Create".
XCMS Online允许您使用树视图从任何现有文件组合轻松创建唯一的数据集。要做到这一点,从顶部导航菜单中选择“Stored Datasets”,点击右上角子菜单中的“Tree View”。你所有的数据集将在屏幕左侧的文件资源管理器中列出。您可以通过单击每个名称左边的复选框来选择整个数据集,或者通过单击三角形来选择部分数据以展开相关文件。一旦您选择了所需的数据,在“数据集名称”字段中为您的新数据集输入一个唯一的名称,然后单击“创建”。
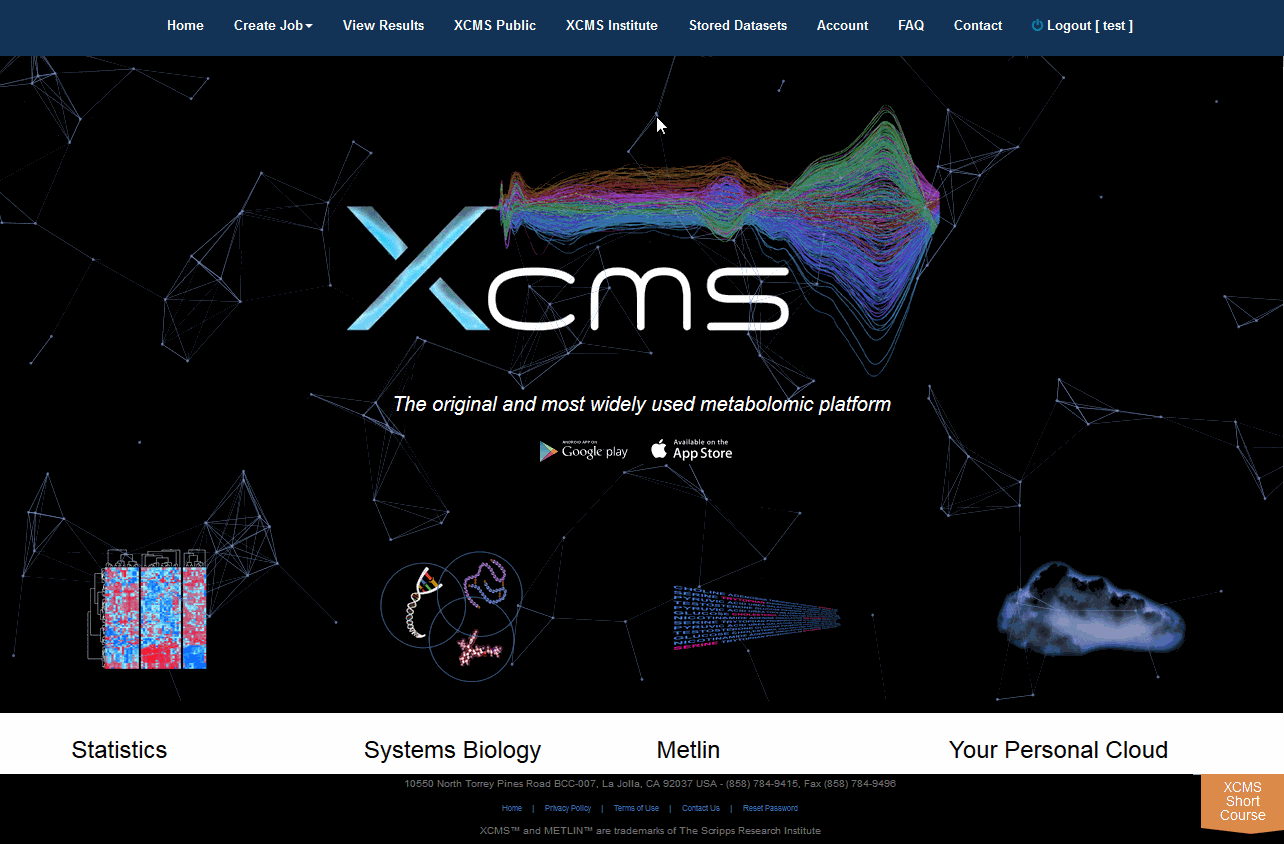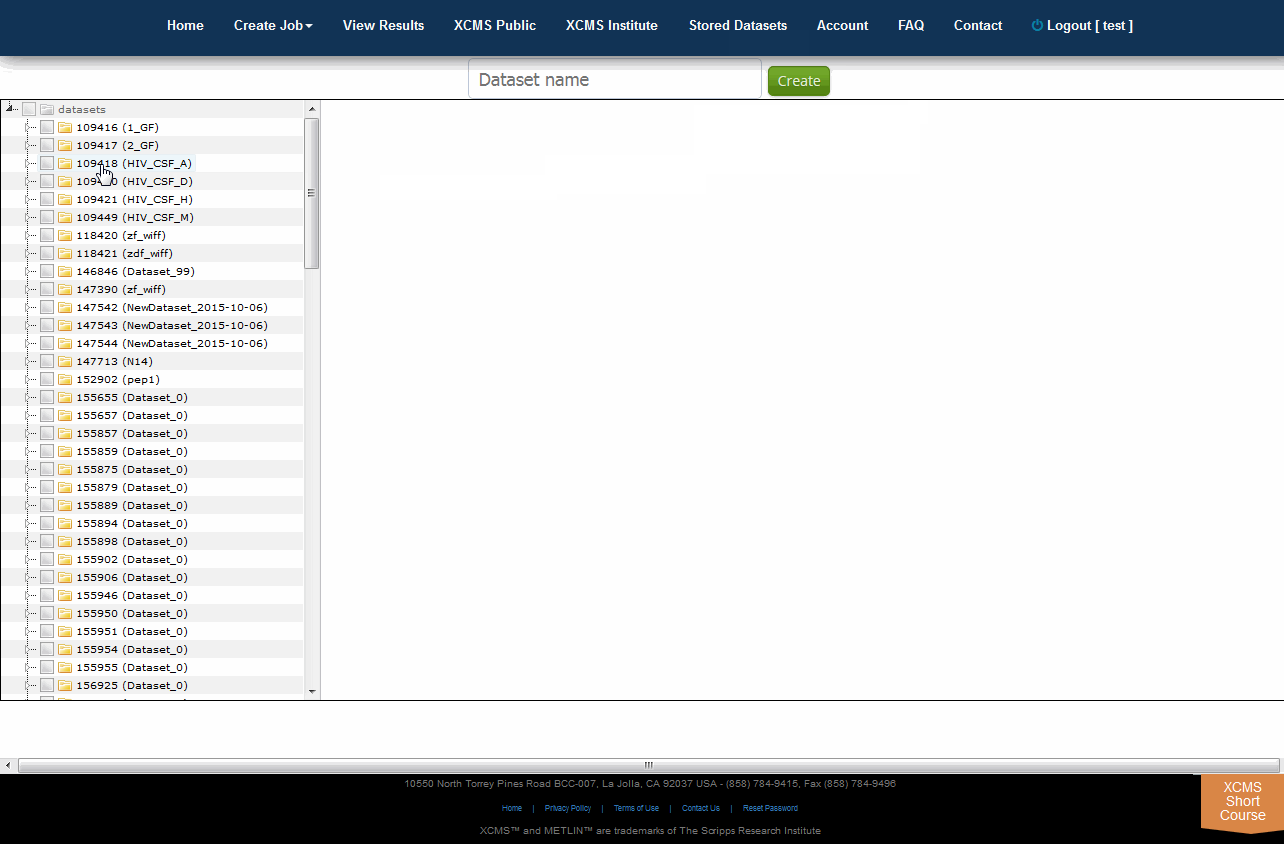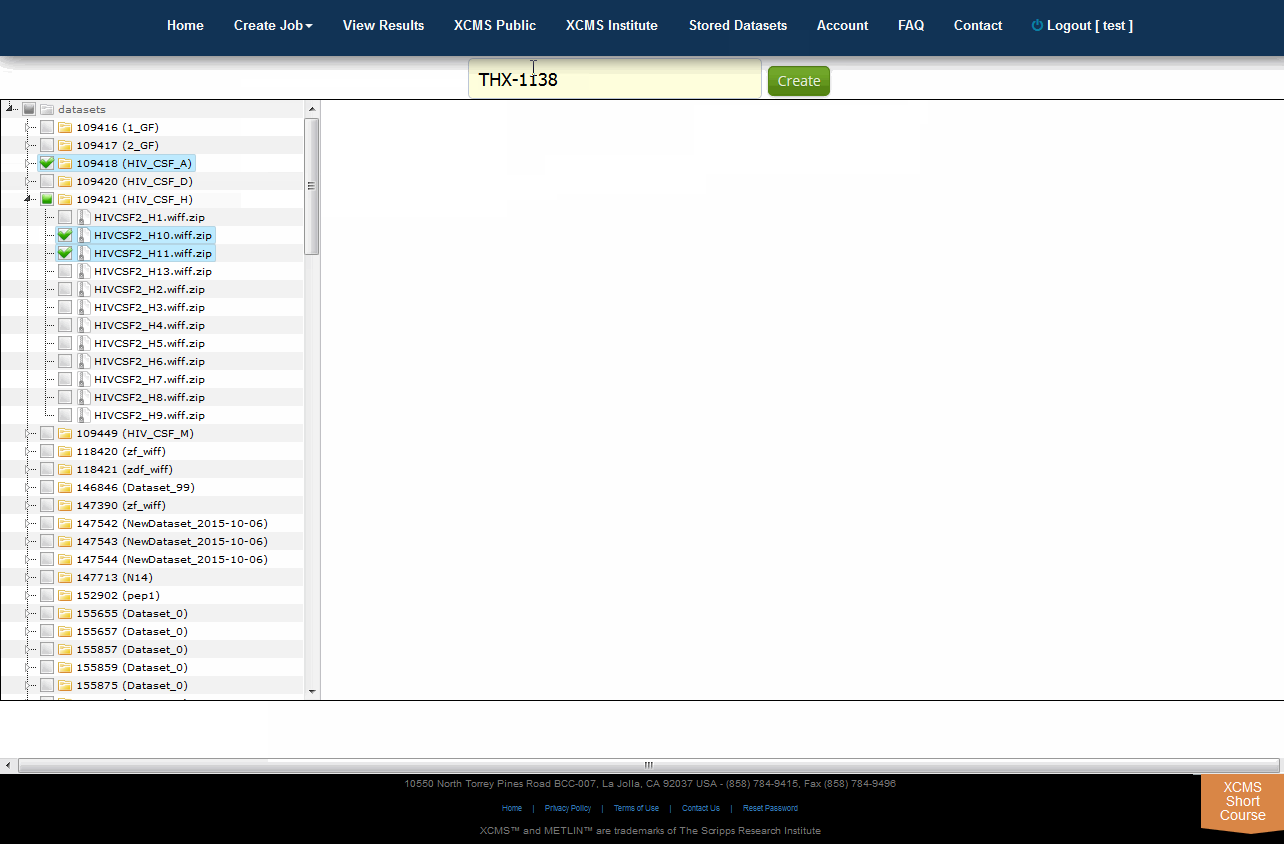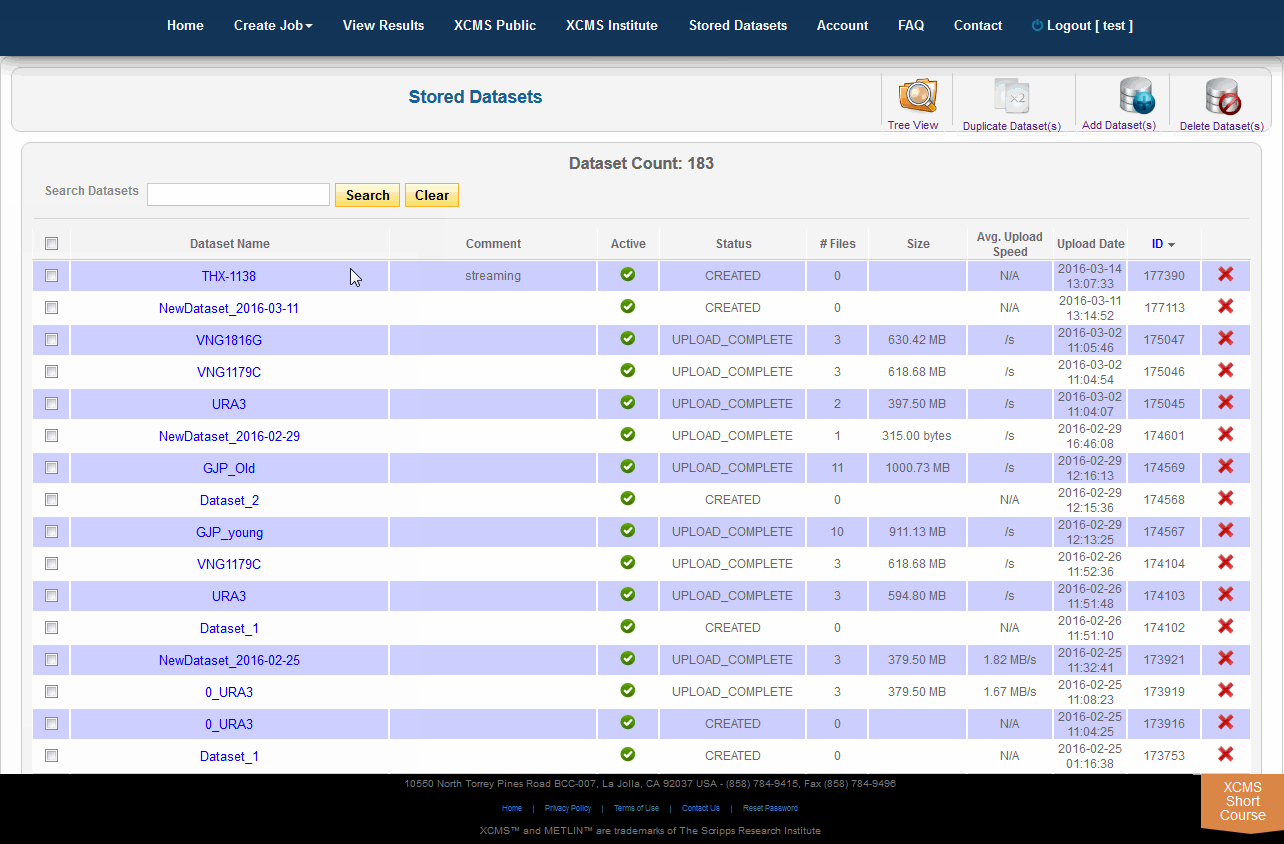
6.0 参数管理
Parameter Management
The Parameter Manager can be accessed via the “Parameter Manager” icon next to “User Settings” in the Account sub menu. Here the you can search, review, create and delete your parameter sets. This will not affect previously-processed jobs unless they are re-run. Resubmitted jobs will use the new parameter set(s) for processing.
可以通过“帐户”子菜单中“用户设置”旁边的“参数管理器”图标访问参数管理器。在这里,您可以搜索、查看、创建和删除参数集。这不会影响以前处理的作业,除非它们被重新运行。重新提交的作业将使用新的参数集进行处理。
7.0 工作共享
Job Sharing
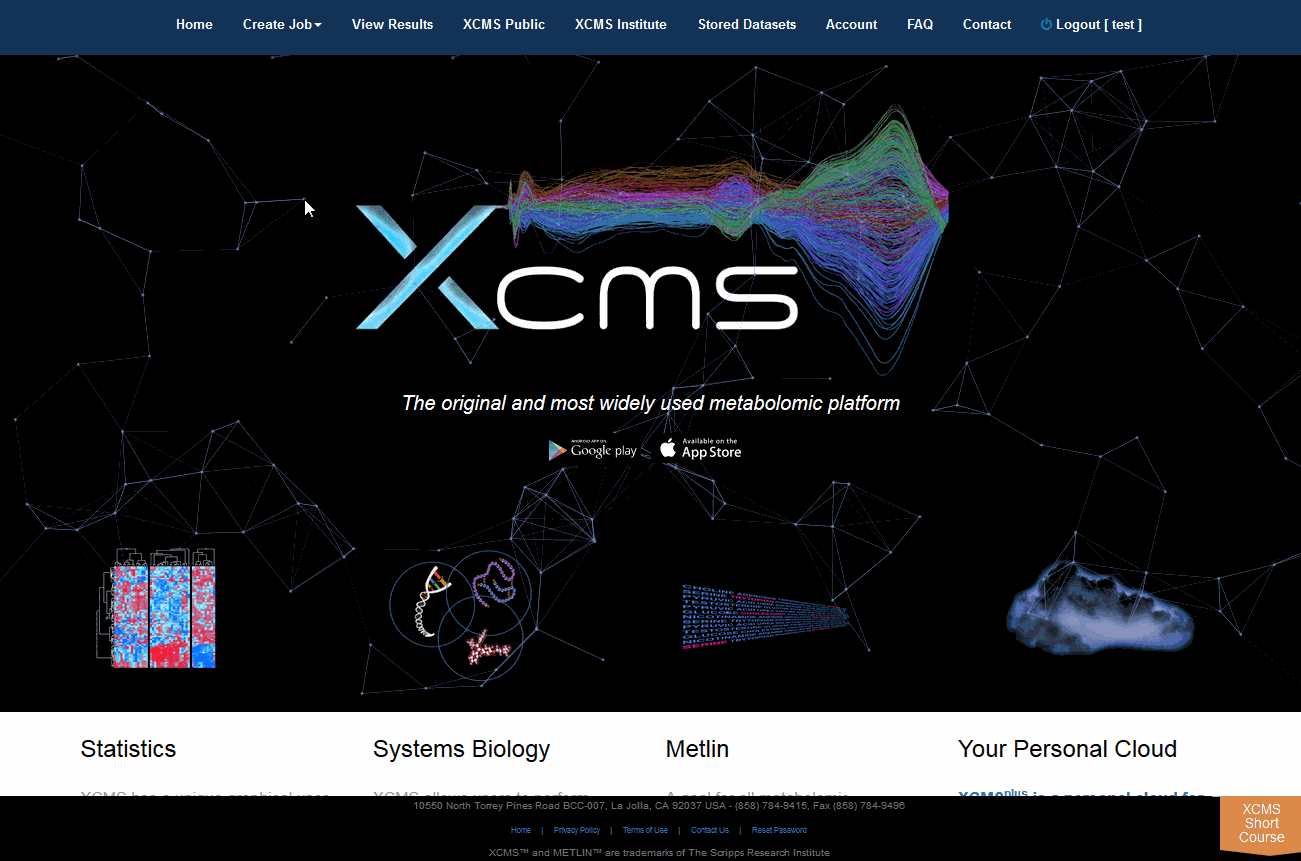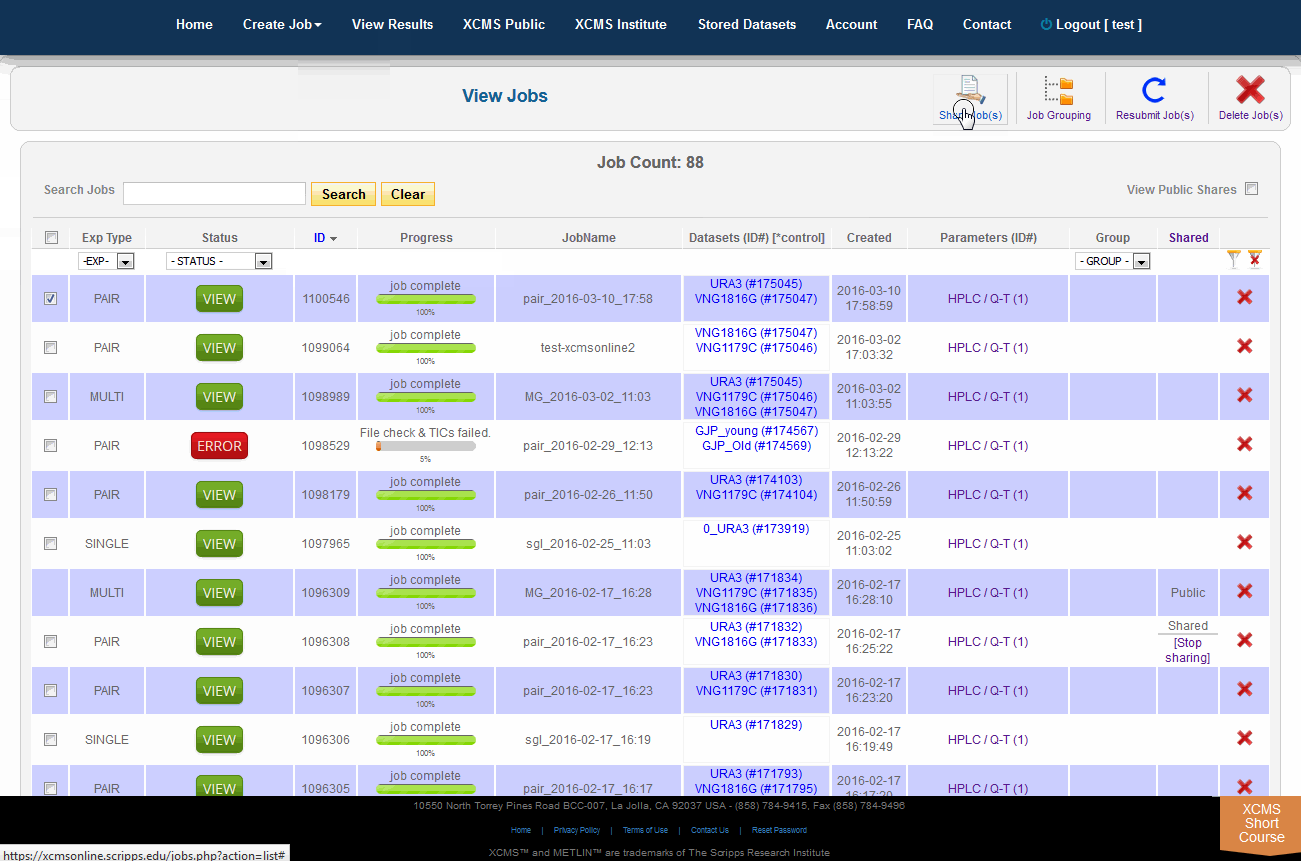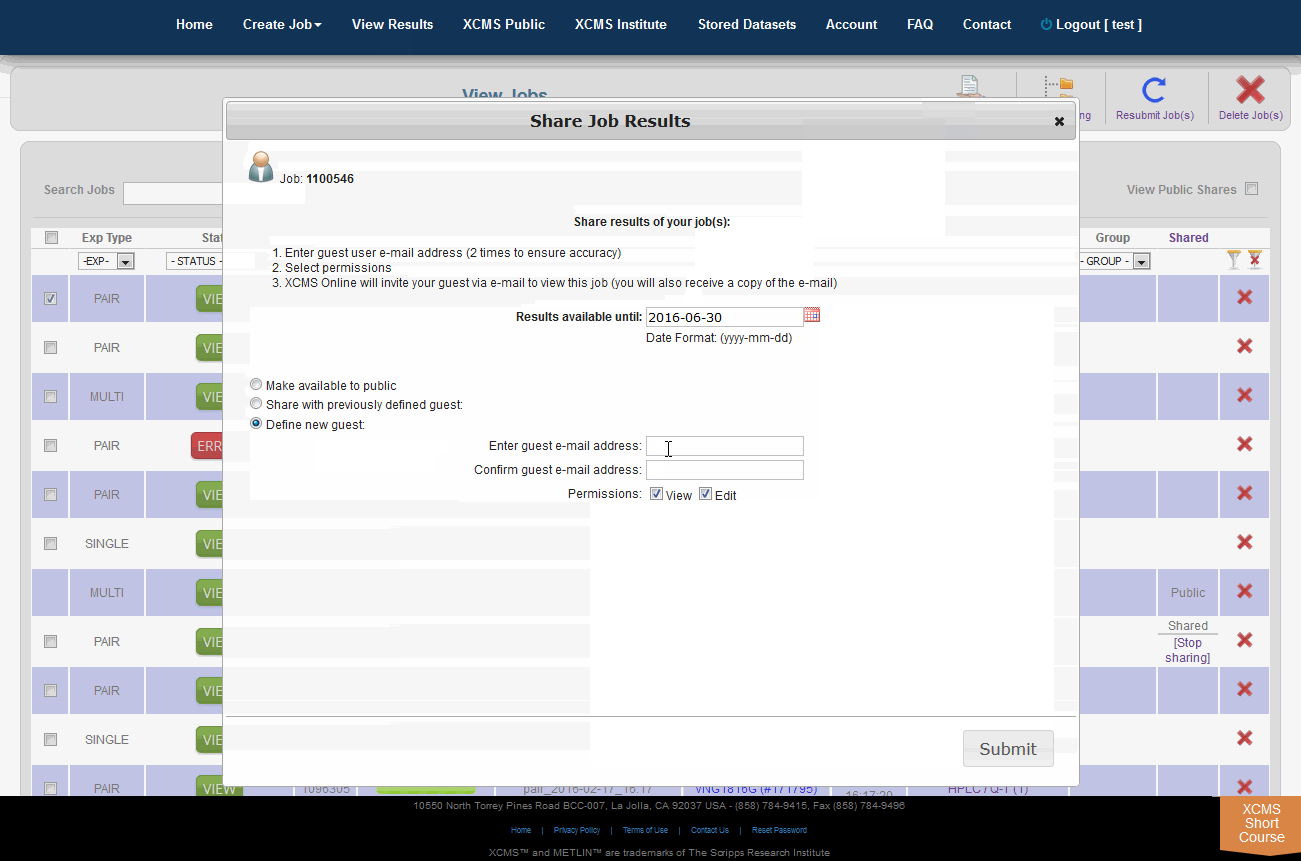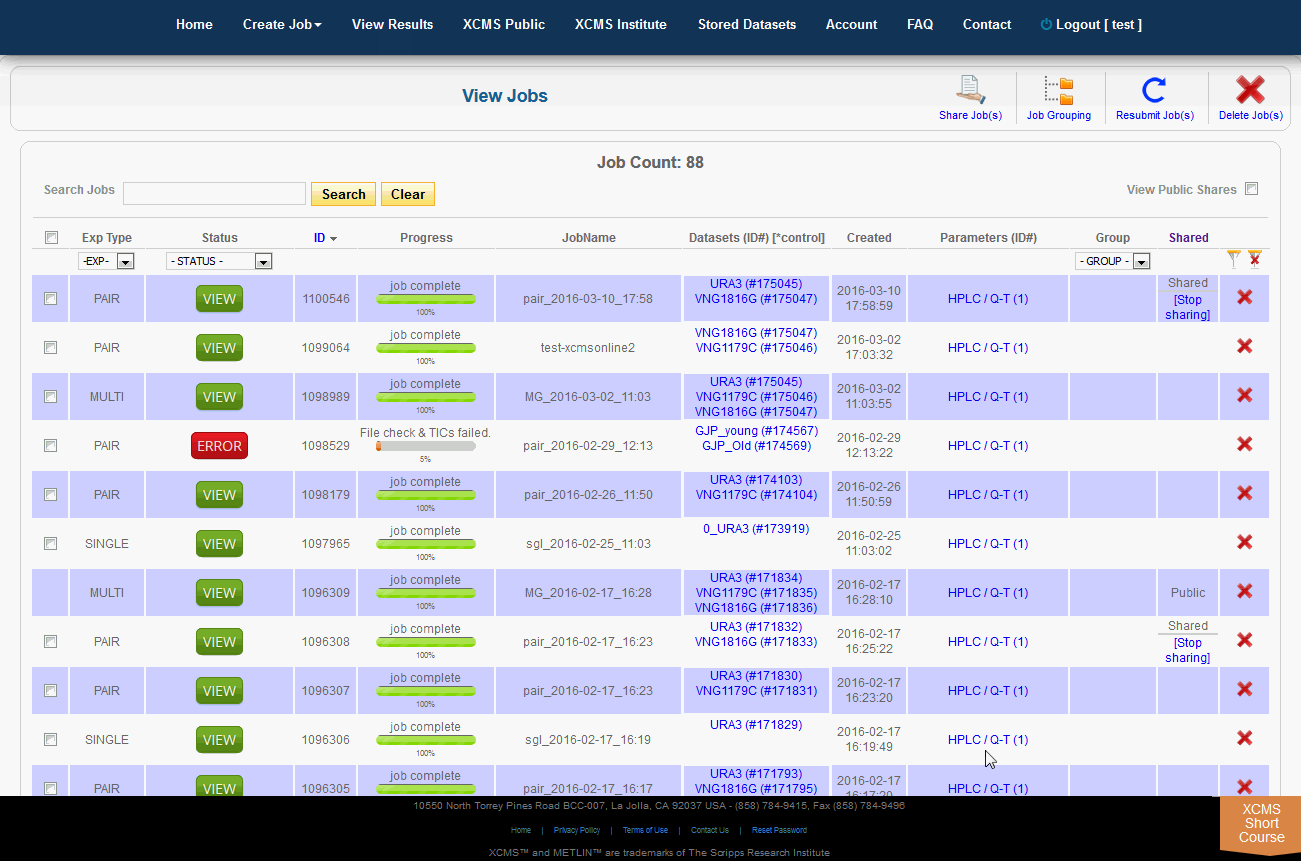
The XCMS Online system allow you to share one job or many jobs instantly with a specific user or all users. From the "View Results" page simply click the check-box to the left of the job you want to share, then click "Share Job(s)" in the upper-right. A pop-up screen will appear.
XCMSOnline系统允许您立即与特定用户或所有用户共享一个作业或多个作业。从“查看结果”页面中,只需单击要共享的作业左边的复选框,然后单击右上角的“共享作业”。将会出现一个弹出式屏幕。
Step 1: Chose how long to share. The default time for a share is three (3) months (unless modified by an administrator). In the field "Results available until" enter the desired expiration date in the format yyyy-mm-dd.
Step 2: Choose who can view the job. There are three options: Share with everyone, share with a previous guest, or create a new guest to share with.
Step 3: Set the permissions. Selecting only the "View" check-box will allow guests to read your data and notes, but they cannot modify it. Selecting the "Edit" check-box allows guests to re-run your job with different parameters and make annotations.
第一步:选择分享的时间。共享的默认时间是三(3)个月(除非管理员修改)。在“Results available until”字段中输入所需的截止日期,格式为“yyyy-mm-dd”。
第二步:选择谁可以查看这份工作。有三种选择:与所有人分享,与以前的客人分享,或创建一个新的客人分享。
第三步:设置权限。只选择“查看”复选框将允许客户读取您的数据和注释,但他们不能修改它。选择“Edit”复选框允许客户使用不同的参数重新运行作业并进行注释。
7.1 查看公开股票
Viewing Public Shares
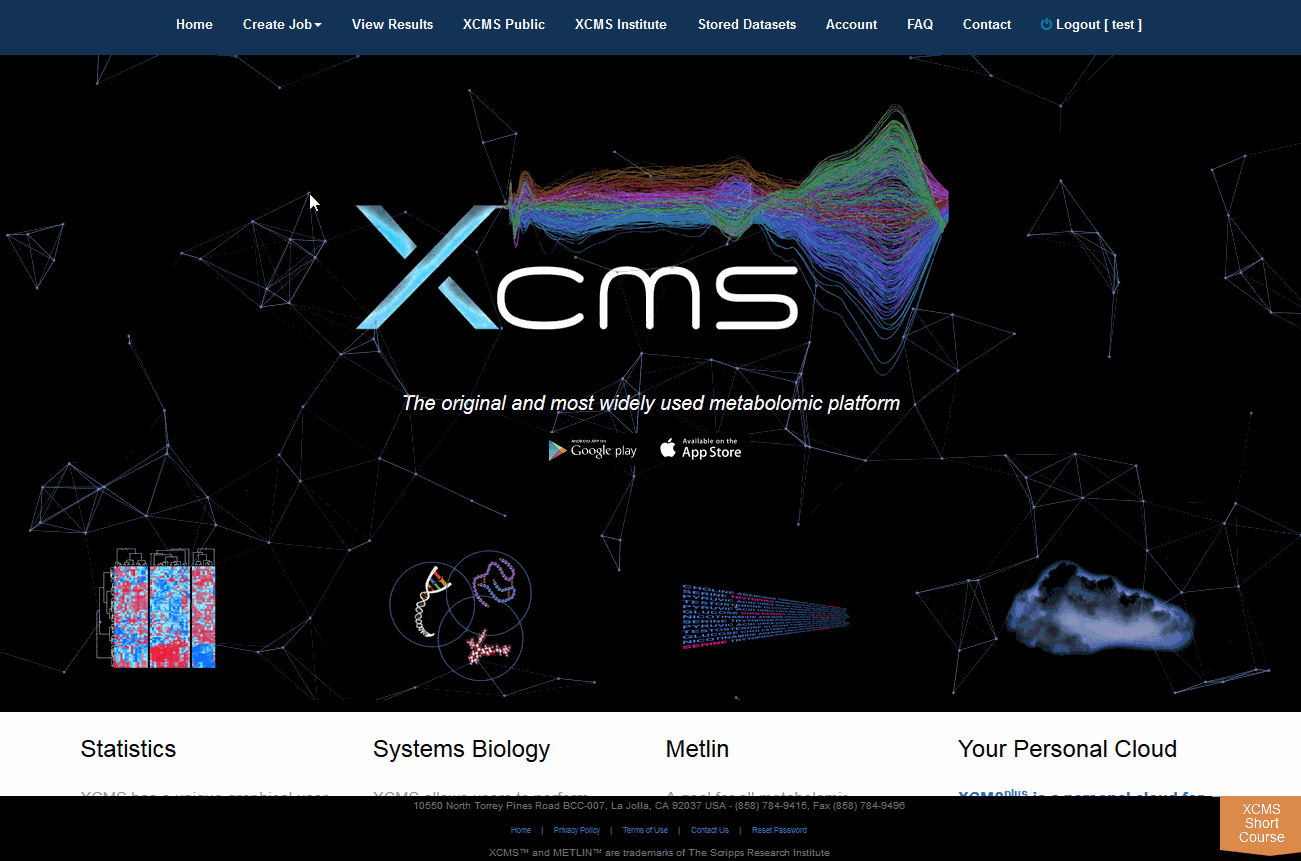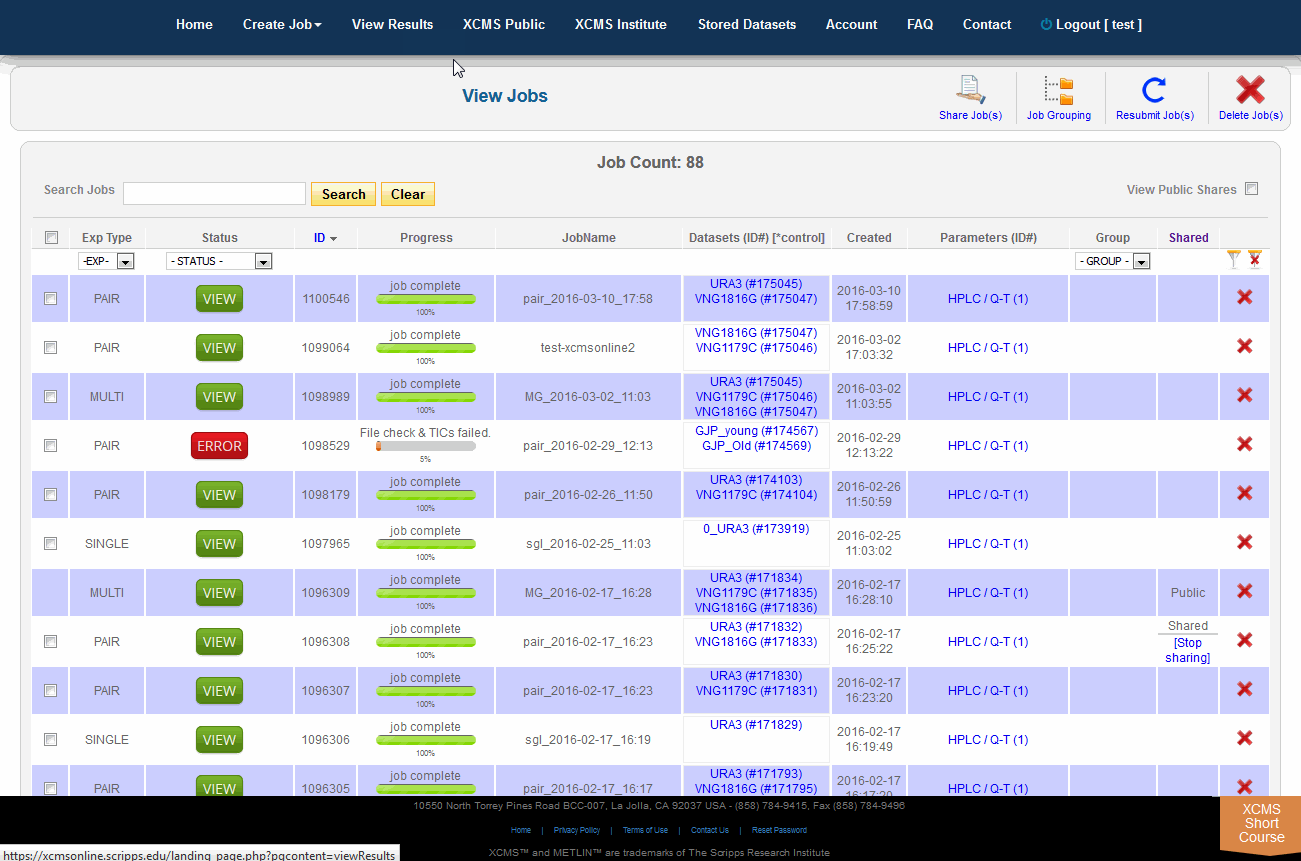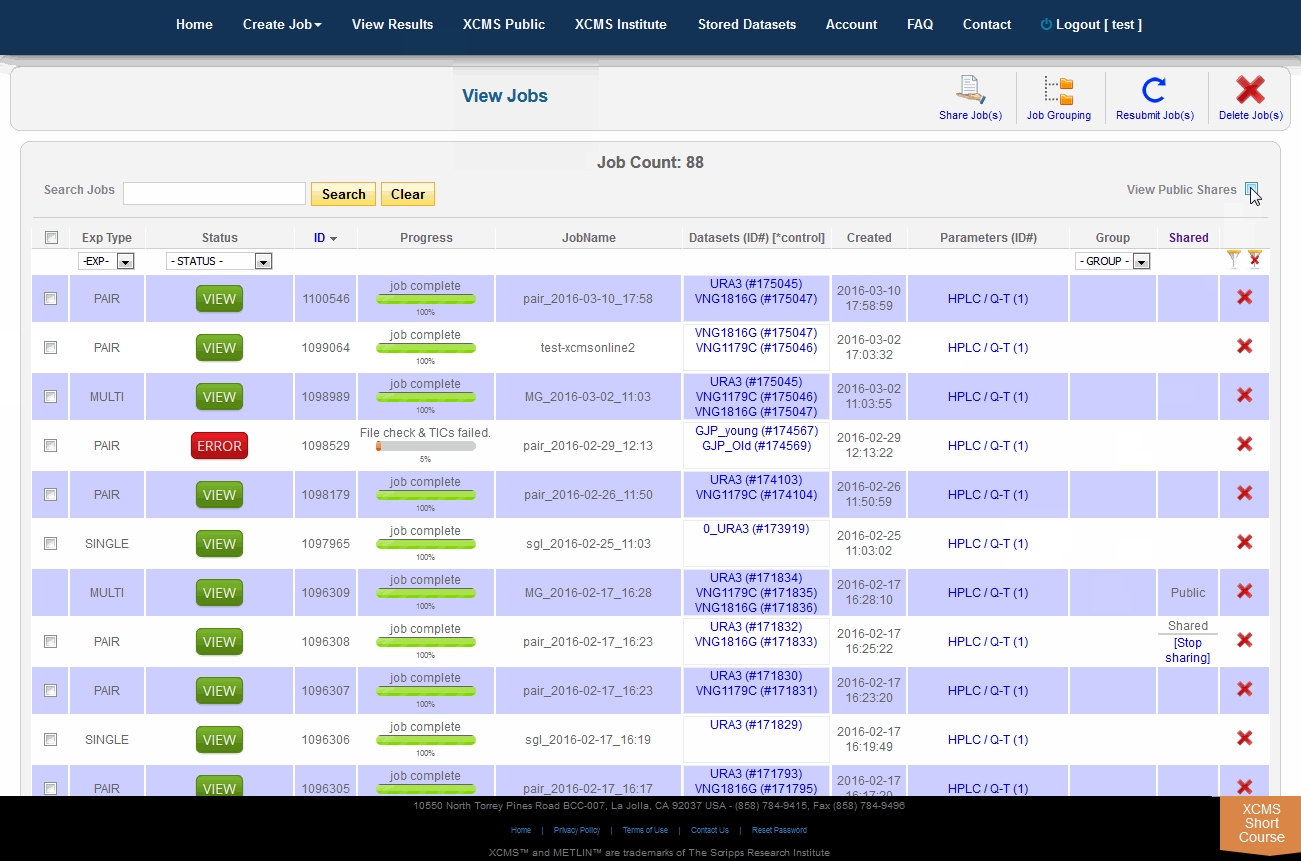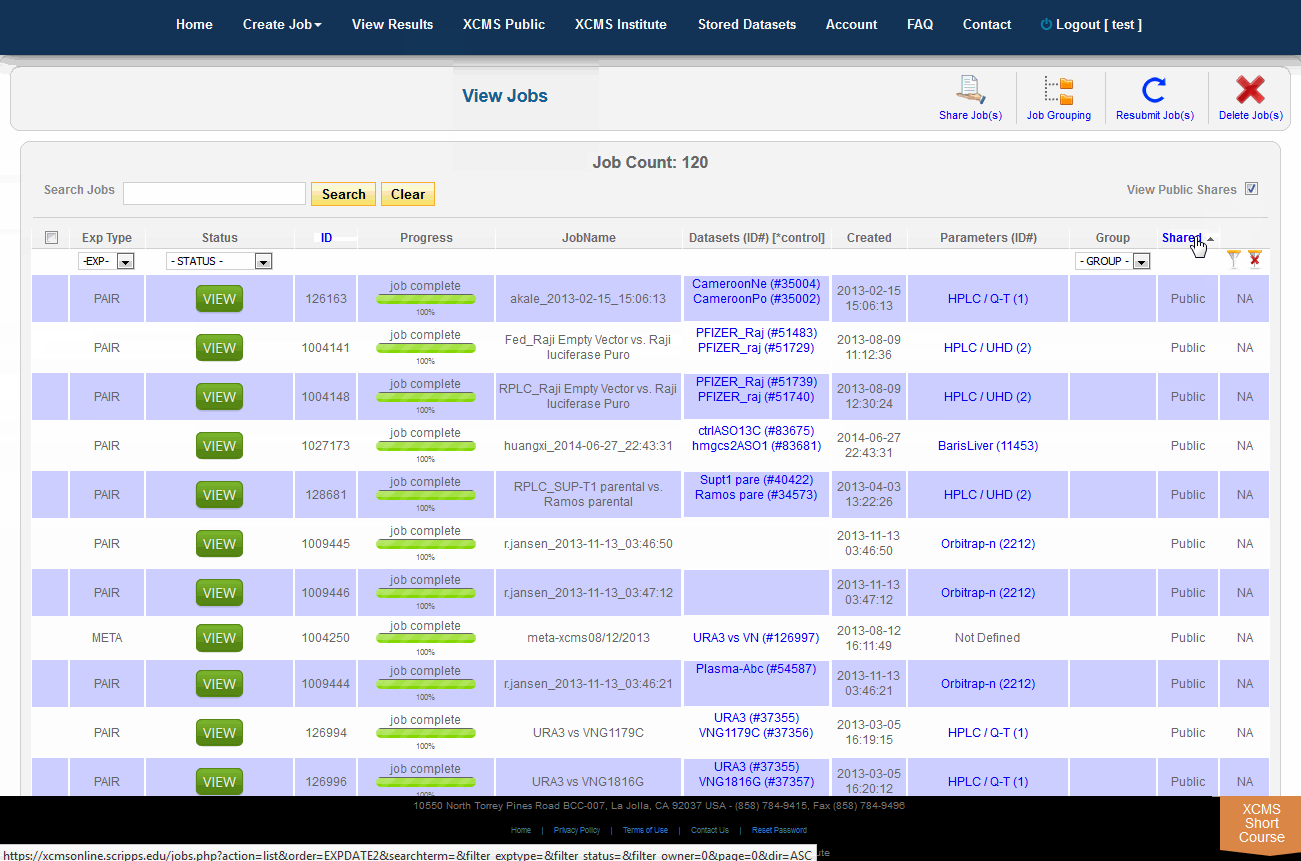
To view job made public by other users, click "View Results" from the top navigation menu. In the upper-right is a check-box named "View Public Shares." After checking the box the page will refresh and include these jobs in the list. To find them easily, click the column heading "Shared" to sort the list by shared and not-shared jobs.
单击顶部导航菜单中的“查看结果”,可查看其他用户公开的作业。右上角有一个名为“View Public Shares”的复选框。选中该框后,页面将刷新并将这些作业包含在列表中。要轻松找到它们,请单击列标题“Shared”,按共享和非共享作业对列表排序。
7.2 共享中心
The Sharing Center
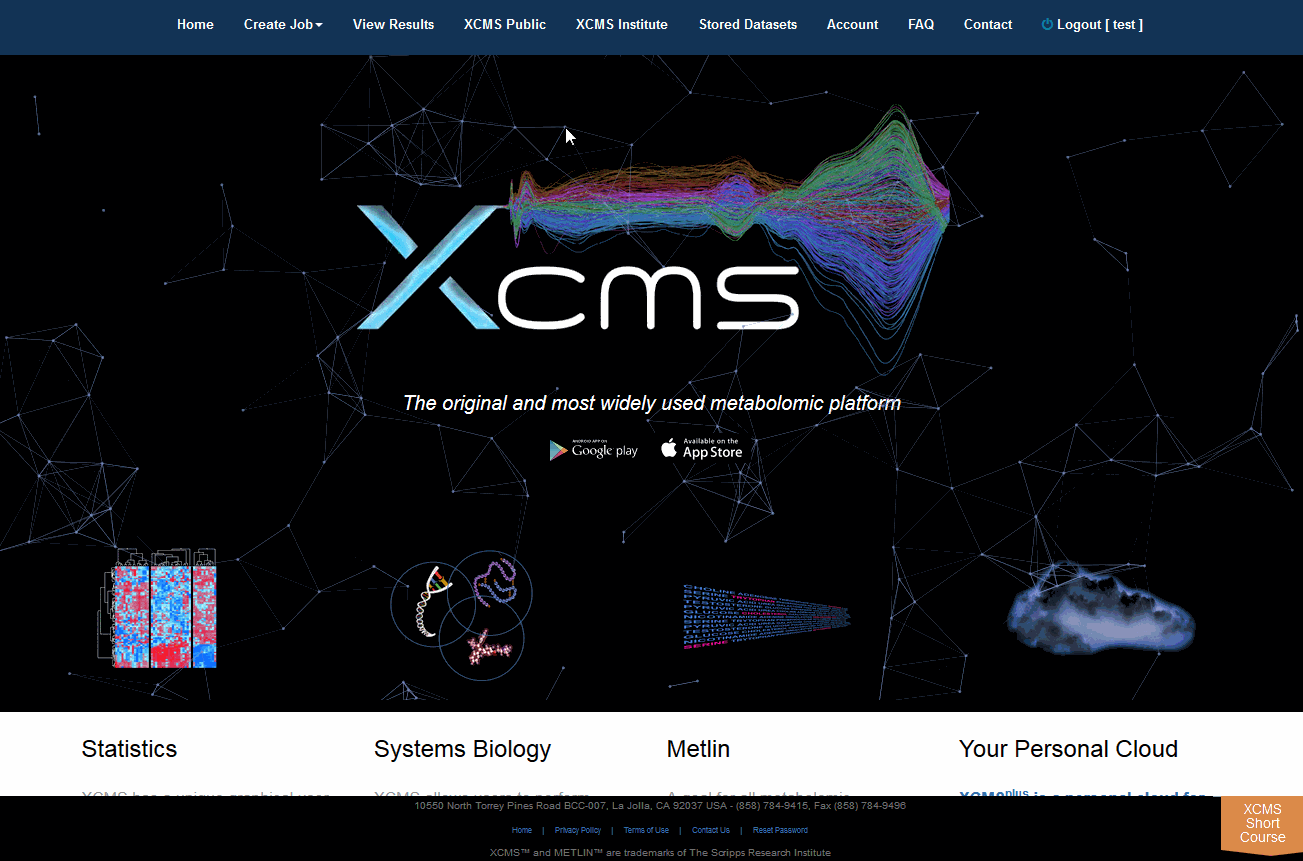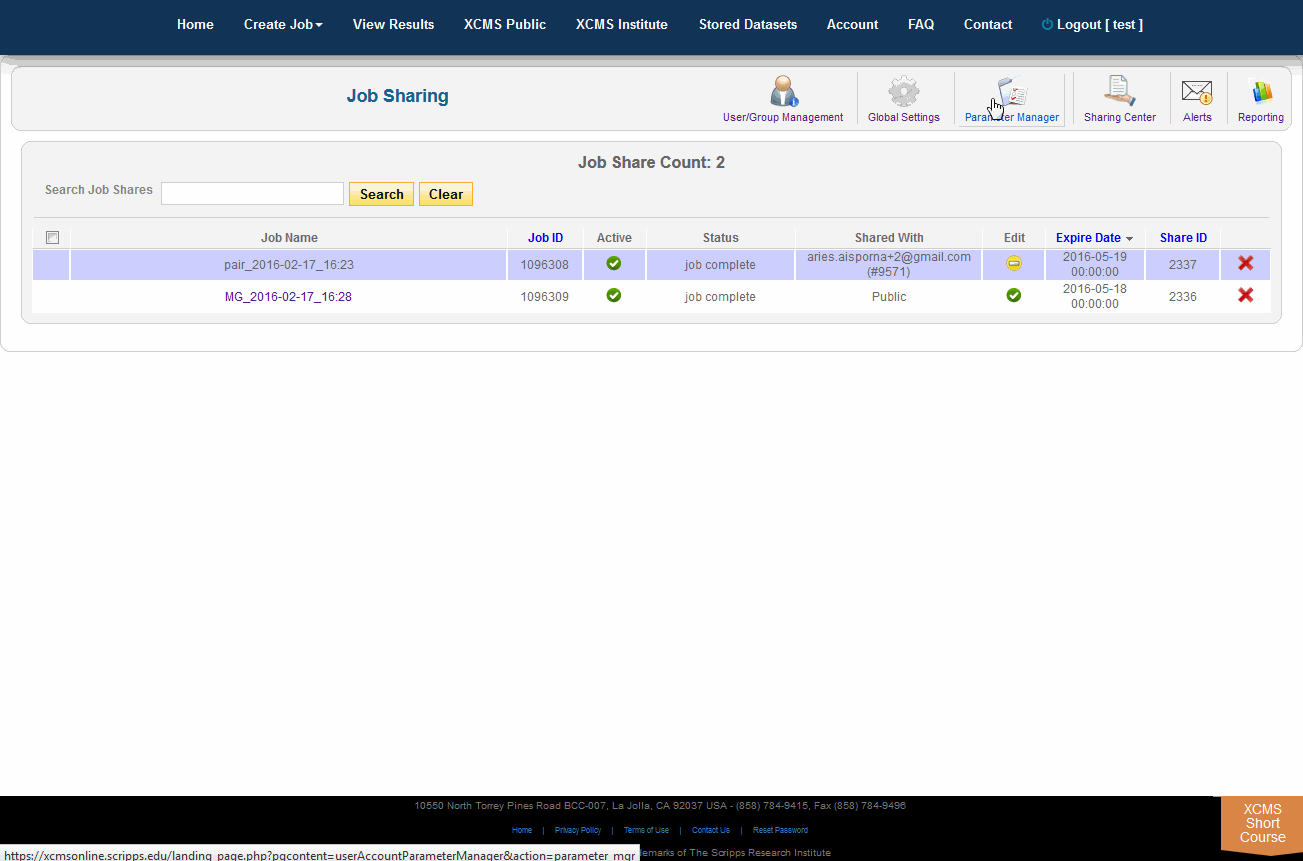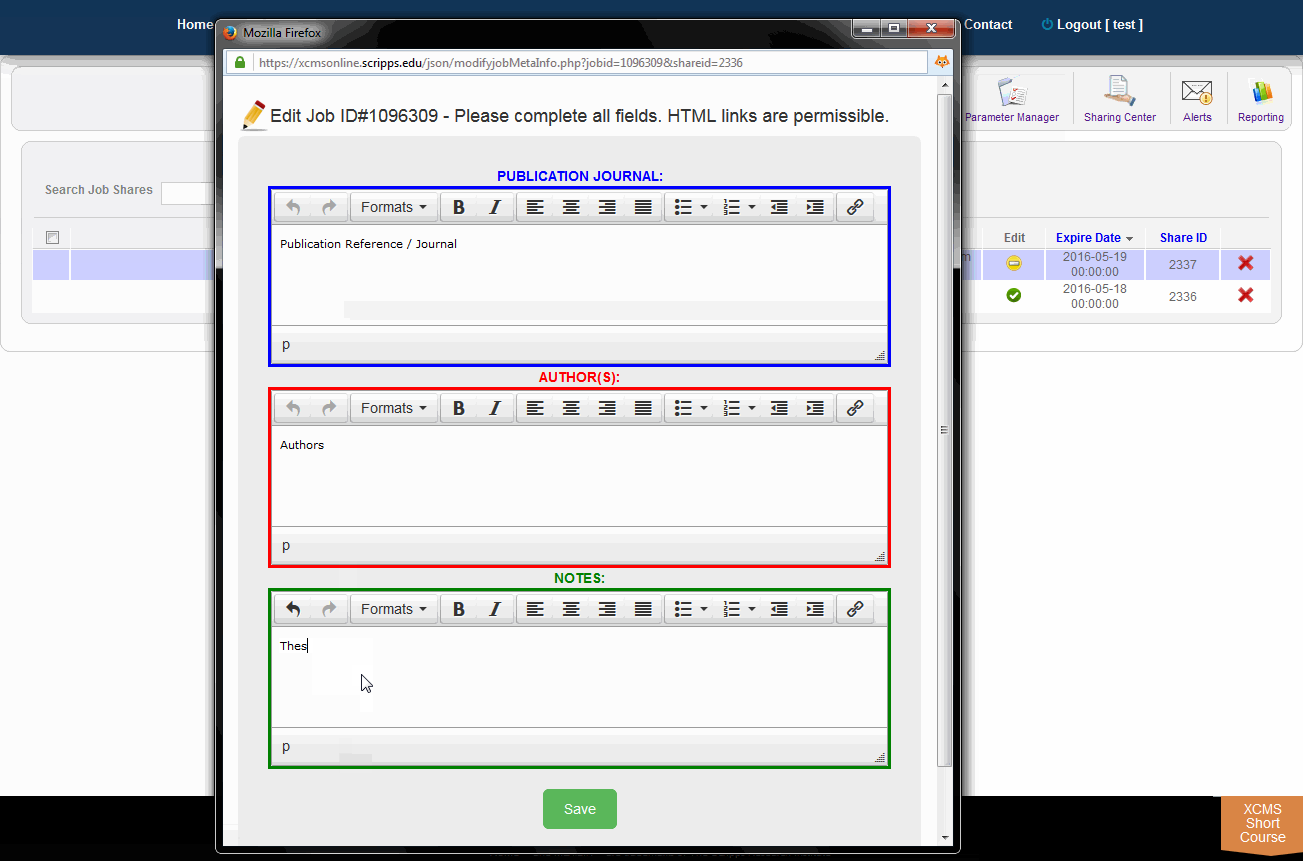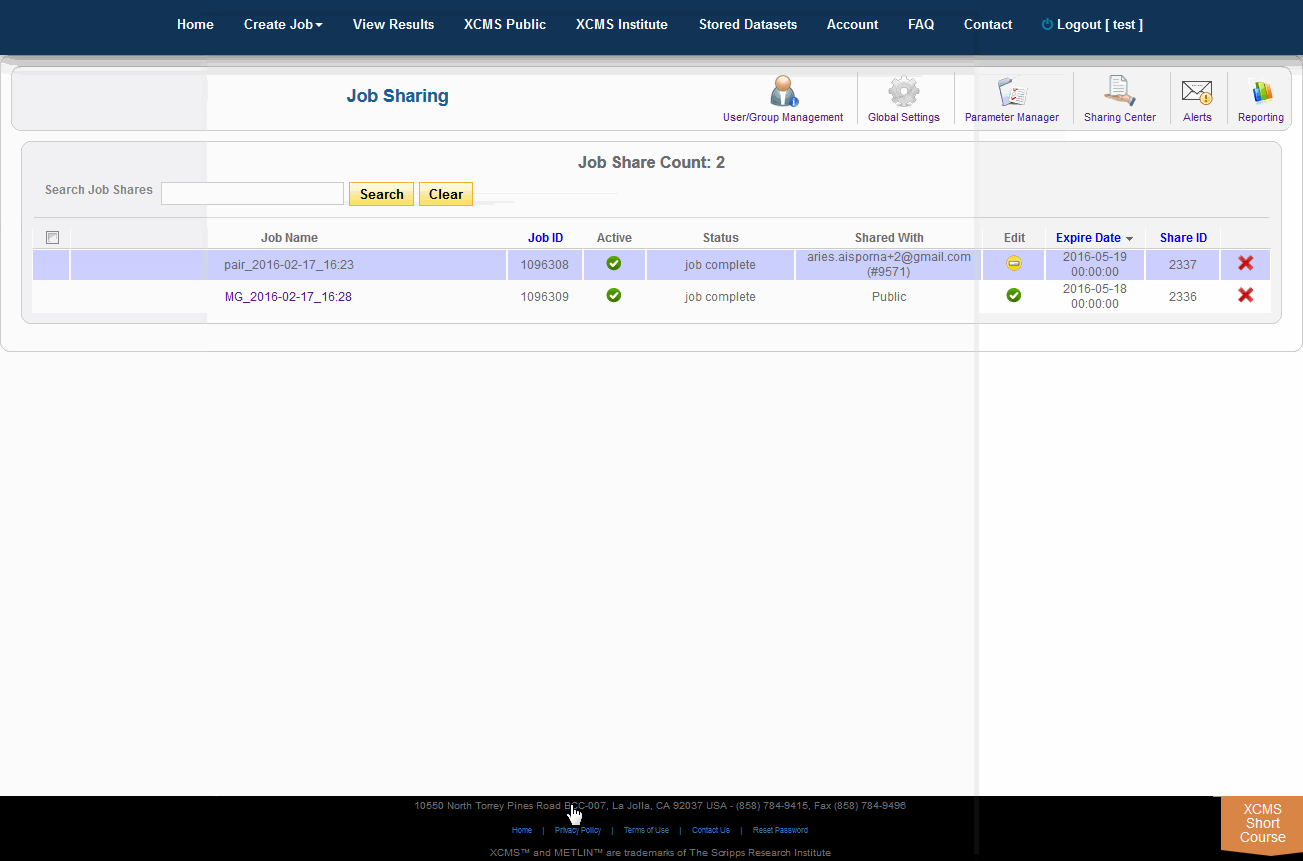
The Sharing Center can be accessed via the “Account” menu in the top navigation bar. Click "Sharing Center". This takes you to a portal where job shares can be viewed, modified and deleted. The page also includes filtering/ordering to see which jobs are scheduled to expire soon. When a job share expires, it will become inaccessible by the guest user, however, the job itself will not be touched. To make the job accessible again the job owner needs to re-share it. Unless otherwise modified by the system administrator the default share period is 3 months.
共享中心可以通过顶部导航栏的“帐户”菜单访问。单击“共享中心”。这将将您带到一个可以查看、修改和删除作业共享的门户。该页还包括筛选/排序,以查看哪些作业计划很快到期。当作业共享到期时,guest用户将无法访问该作业,但是作业本身不会被修改。要使作业再次可访问,作业所有者需要重新共享它。除非系统管理员另有修改,否则默认的共享期为3个月。
You can add annotations to public shares by clicking the job name. A window will pop-up with the fields "Publication Journal," "Authors," and "Notes." Make your changes to these fields and click the green "Save" button.
您可以通过单击作业名称向公共共享添加注释。将弹出一个窗口,其中包含“Publication Journal”、“Authors”和“Notes”字段。对这些字段进行更改,然后单击绿色的“保存”按钮。